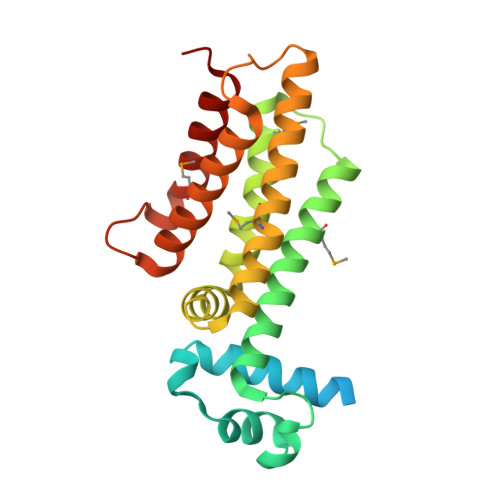
Structure of EthR in a Ligand Bound Conformation Reveals Therapeutic Perspectives against Tuberculosis
Frenois, F., Engohang-Ndong, J., Locht, C., Baulard, A.R., Villeret, V.(2004) Mol Cell 16: 301-307
- PubMed: 15494316 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.09.020
- Primary Citation Related Structures:
1U9N, 1U9O - PubMed Abstract:
Mycobacterium tuberculosis EthR is a repressor of ethA, a gene encoding a mono-oxygenase required for the activation of the prodrug ethionamide. Here we describe the X-ray crystal structure of EthR, a homodimer with an entirely helical structure showing similarities to TetR family members. Each monomer contained a fortuitous ligand identified as hexadecyl octanoate. The crystal structure of EthR purified in M. smegmatis revealed the presence of a comparable ligand. The binding of hexadecyl octanoate to EthR induces a conformational state incompatible with repressor function, which should lead to ethA derepression and consequently to an increased sensitivity to ethionamide and other thioamides. A related, more hydrophilic ketone was found to exhibit synergistic antimycobacterial effects when tested together with ethionamide, indicating that this strategy may help reduce the dosage of potent antibacterial compounds that otherwise are too toxic to be used as first-line drugs.
- CNRS-UMR8525, 1 rue du Professeur Calmette, BP245 59019 Lille cedex, France.
Organizational Affiliation: