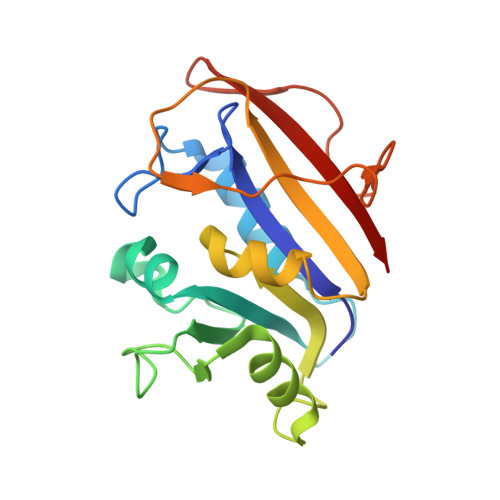
Understanding the role of Leu22 variants in methotrexate resistance: comparison of wild-type and Leu22Arg variant mouse and human dihydrofolate reductase ternary crystal complexes with methotrexate and NADPH.
Cody, V., Luft, J.R., Pangborn, W.(2005) Acta Crystallogr D Biol Crystallogr 61: 147-155
- PubMed: 15681865 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904030422
- Primary Citation Related Structures:
1U70, 1U71, 1U72 - PubMed Abstract:
Structural data are reported to 2.5 A resolution for the first full analysis of the methotrexate-resistant Leu22Arg (L22R) variant of mouse dihydrofolate reductase (mDHFR) crystallized as a ternary complex with methotrexate (MTX) and the cofactor NADPH. These results are compared with the MTX and NADPH ternary complexes of L22R human DHFR (hDHFR) and those of mouse and human wild-type DHFR enzymes. The conformation of mDHFR Arg22 is such that it makes hydrogen-bonding contacts with Asp21, Trp24 and a structural water molecule, observations which were not made in the L22R hDHFR ternary complex. These data show that there is little difference between the structures of the wild type and L22R variant for either mouse or human DHFR; however, there are significant differences between the species. Comparison of these structures reveals that the active site of mDHFR is larger than that in the hDHFR structure. In mDHFR, the position of MTX is shifted 0.6 A toward helix C (residues 59-65), which in turn is shifted 1.2 A away from the active site relative to that observed in the hDHFR ternary complexes. In the L22R variant mDHFR structure, MTX makes shorter contacts to the conserved residues Ile7, Val115 and Tyr121 than in the L22R variant human DHFR structure. These contacts are comparable in both wild-type enzymes. The unexpected results from this comparison of the mouse and human DHFR complexes bound with the same ligand and cofactor illustrate the importance of detailed study of several species of enzyme, even when there is a high sequence homology between them. These data suggest that the differences in binding interactions of the L22R variant are in agreement with the weaker binding affinity for MTX in the variant enzymes; the larger size of the binding site in mDHFR supports the observation that the binding affinity of MTX for L22R mDHFR is significantly weaker than that of the L22R hDHFR enzyme.
- Hauptman-Woodward Medical Research Institute, 73 High Street, Buffalo, NY 14203, USA. cody@hwi.buffalo.edu
Organizational Affiliation: