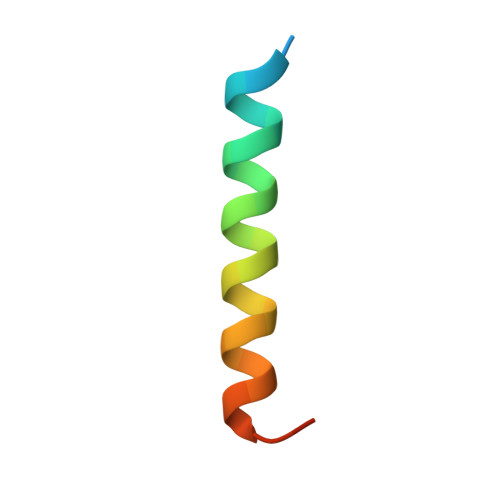
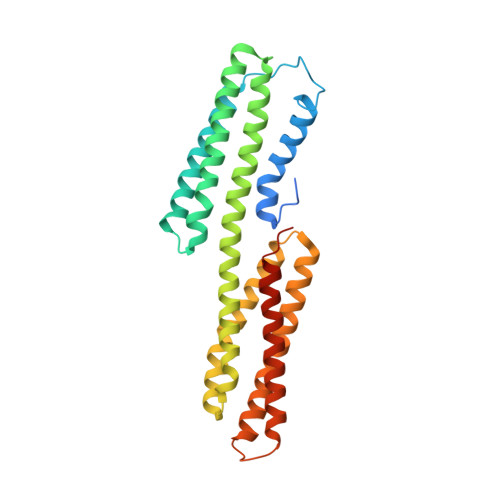
A vinculin binding domain from the talin rod unfolds to form a complex with the vinculin head.
Fillingham, I., Gingras, A.R., Papagrigoriou, E., Patel, B., Emsley, J., Critchley, D.R., Roberts, G.C., Barsukov, I.L.(2005) Structure 13: 65-74
- PubMed: 15642262 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.11.006
- Primary Citation Related Structures:
1U6H, 1U89 - PubMed Abstract:
The cytoskeletal protein talin plays a key role in activating integrins and in coupling them to the actin cytoskeleton. Its N-terminal globular head, which binds beta integrins, is linked to an extended rod having a C-terminal actin binding site and several vinculin binding sites (VBSs). The NMR structure of residues 755-889 of the rod (containing a VBS) is shown to be an amphipathic four-helix bundle with a left-handed topology. A talin peptide corresponding to the VBS binds the vinculin head; the X-ray crystallographic structure of this complex shows that the residues which interact with vinculin are buried in the hydrophobic core of the talin fragment. NMR shows that the interaction involves a major structural change in the talin fragment, including unfolding of one of its helices, making the VBS accessible to vinculin. Interestingly, the talin 755-889 fragment binds more than one vinculin head molecule, suggesting that the talin rod may contain additional as yet unrecognized VBSs.
- Department of Biochemistry, University of Leicester, Leicester LE1 7RH, UK.
Organizational Affiliation: