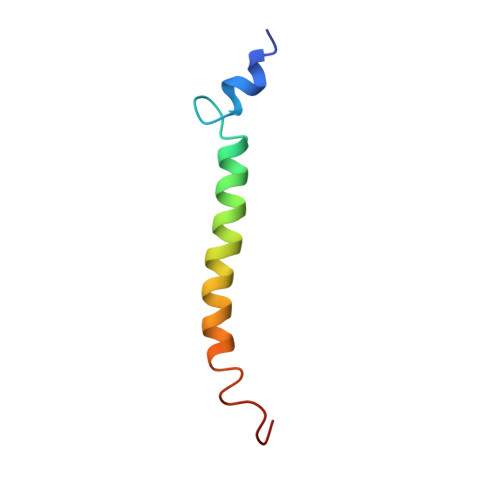
Helical structure determined by NMR of the HIV-1 (345-392)Gag sequence, surrounding p2: Implications for particle assembly and RNA packaging
Morellet, N., Druillennec, S., Lenoir, C., Bouaziz, S., Roques, B.P.(2005) Protein Sci 14: 375-386
- PubMed: 15659370
- DOI: https://doi.org/10.1110/ps.041087605
- Primary Citation Related Structures:
1U57 - PubMed Abstract:
Gag protein oligomerization, an essential step during virus assembly, results in budding of spherical virus particles. This process is critically dependent on the spacer p2, located between the capsid and the nucleocapsid proteins. P2 contributes also, in association with NCp7, to specific recognition of the HIV-1 packaging signal resulting in viral genome encapsidation. There is no structural information about the 20 last amino acids of the C-terminal part of capsid (CA[CTD]) and p2, in the molecular mechanism of Gag assembly. In this study the structure of a peptide encompassing the 14 residues of p2 with the upstream 21 residues and the downstream 13 residues was determined by (1)H NMR in 30% trifluoroethanol (TFE). The main structural motif is a well-defined amphipathic alpha-helix including p2, the seven last residues of the CA(CTD), and the two first residues of NCp7. Peptides containing the p2 domain have a strong tendency to aggregate in solution, as shown by gel filtration analyses in pure H(2)O. To take into account the aggregation phenomena, models of dimer and trimer formed through hydrophobic or hydrophilic interfaces were constructed by molecular dynamic simulations. Gel shift experiments demonstrate that the presence of at least p2 and the 13 first residues of NCp7 is required for RNA binding. A computer-generated model of the Gag polyprotein segment (282-434)Gag interacting with the packaging element SL3 is proposed, illustrating the importance of p2 and NCp7 in genomic encapsidation.
- Unite de Pharmacologie Chimique et Genetique, INSERM U640, CNRS UMR 8151, UFR des Sciences Pharmaceutiques et Biologiques, 4, Avenue de l'Observatoire, 75270 Paris Cedex 06, France. morellet@pharmacie.univ-paris5.fr
Organizational Affiliation: