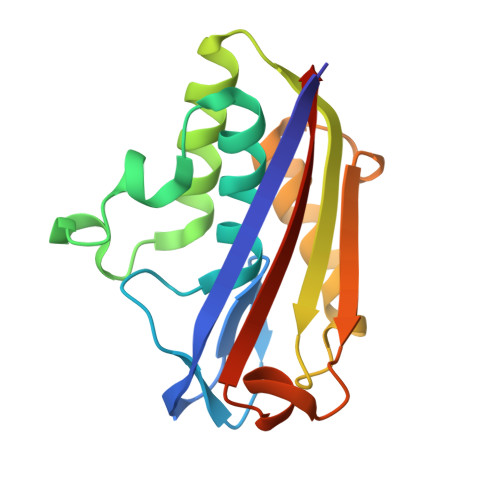
Structure of 2C-Methyl-D-Erythritol-2,4-Cyclodiphosphate Synthase Involved in Mevalonate Independent Biosynthesis of Isoprenoids
Steinbacher, S., Kaiser, J., Wungsintaweekul, J., Hecht, S., Eisenreich, W., Gerhardt, S., Bacher, A., Rohdich, F.(2002) J Mol Biology 316: 79-88
- PubMed: 11829504 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5341
- Primary Citation Related Structures:
1JY8, 1U3L, 1U3P, 1U40, 1U43 - PubMed Abstract:
Isoprenoids are biosynthesized from isopentenyl diphosphate and the isomeric dimethylallyl diphosphate via the mevalonate pathway or a mevalonate-independent pathway that was identified during the last decade. The non-mevalonate pathway is present in many bacteria, some algae and in certain protozoa such as the malaria parasite Plasmodium falciparum and in the plastids of higher plants, but not in mammals and archaea. Therefore, these enzymes have been recognised as promising drug targets. We report the crystal structure of Escherichia coli 2C- methyl-d-erythritol-2,4-cyclodiphosphate synthase (IspF), which converts 4-diphosphocytidyl-2C-methyl-d-erythritol 2-phosphate into 2C-methyl-d-erythritol 2,4-cyclodiphosphate and CMP in a Mg-dependent reaction. The protein forms homotrimers that tightly bind one zinc ion per subunit at the active site, which helps to position the substrate for direct attack of the 2-phosphate group on the beta-phosphate.
- Abteilung für Strukturforschung, Max-Planck-Institut für Biochemie, Am Klopferspitz 18a, Martinsried, D-82152, Germany. steinbac@biochem.mpg.de
Organizational Affiliation: