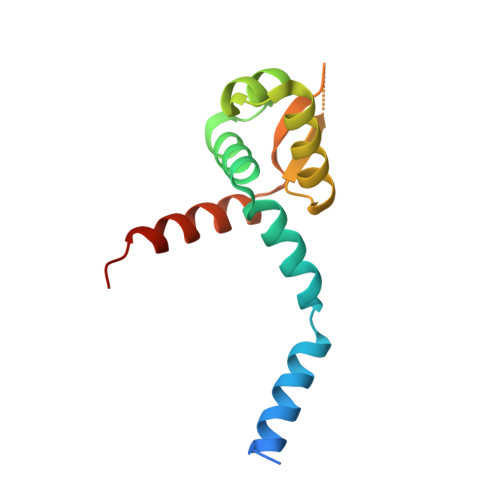
Crystal structure of the Staphylococcus aureus pI258 CadC Cd(II)/Pb(II)/Zn(II)-responsive repressor
Ye, J., Kandegedara, A., Martin, P., Rosen, B.P.(2005) J Bacteriol 187: 4214-4221
- PubMed: 15937183 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.187.12.4214-4221.2005
- Primary Citation Related Structures:
1U2W - PubMed Abstract:
The Staphylococcus aureus plasmid pI258 cadCA operon encodes a P-type ATPase, CadA, that confers resistance to the heavy metals Cd(II), Zn(II), and Pb(II). Expression of this heavy-metal efflux pump is regulated by CadC, a homodimeric repressor that dissociates from the cad operator/promoter upon binding of Cd(II), Pb(II), or Zn(II). CadC is a member of the ArsR/SmtB family of metalloregulatory proteins. Here we report the X-ray crystal structure of CadC at 1.9 angstroms resolution. The dimensions of the protein dimer are approximately 30 angstroms by 40 angstroms by 70 angstroms. Each monomer contains six alpha-helices and a three-stranded beta-sheet. Helices 4 and 5 form a classic helix-turn-helix motif that is the putative DNA binding region. The alpha1 helix of one monomer crosses the dimer to approach the alpha4 helix of the other monomer, consistent with the previous proposal that these two regulatory metal binding sites for the inducer cadmium or lead are each formed by Cys-7 and Cys-11 from the N terminus of one monomer and Cys-58 and Cys-60 of the other monomer. Two nonregulatory metal binding sites containing zinc are formed between the two antiparallel alpha6 helices at the dimerization interface. This is the first reported three-dimensional structure of a member of the ArsR/SmtB family with regulatory metal binding sites at the DNA binding domain and the first structure of a transcription repressor that responds to the heavy metals Cd(II) and Pb(II).
- Department of Biochemistry and Molecular Biology, Wayne State University, School of Medicine, 540 E. Canfield Ave., Detroit, MI 48201, USA. .
Organizational Affiliation: