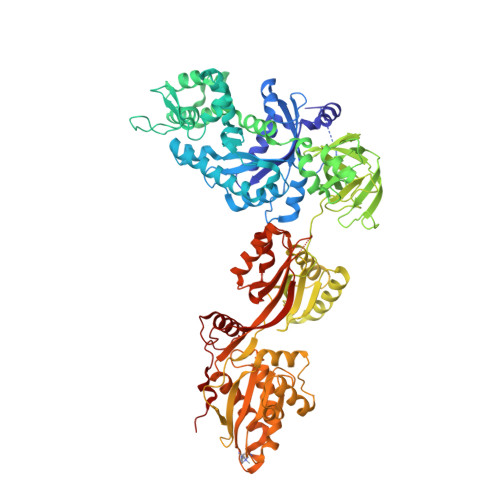
Crystal Structure of ADP-ribosylated Ribosomal Translocase from Saccharomyces cerevisiae
Jorgensen, R., Yates, S.P., Teal, D.J., Nilsson, J., Prentice, G.A., Merrill, A.R., Andersen, G.R.(2004) J Biological Chem 279: 45919-45925
- PubMed: 15316019
- DOI: https://doi.org/10.1074/jbc.M406218200
- Primary Citation Related Structures:
1U2R - PubMed Abstract:
The crystal structure of ADP-ribosylated yeast elongation factor 2 in the presence of sordarin and GDP has been determined at 2.6 A resolution. The diphthamide at the tip of domain IV, which is the target for diphtheria toxin and Pseudomonas aeruginosa exotoxin A, contains a covalently attached ADP-ribose that functions as a very potent inhibitor of the factor. We have obtained an electron density map of ADP-ribosylated translation factor 2 revealing both the ADP-ribosylation and the diphthamide. This is the first structure showing the conformation of an ADP-ribosylated residue and confirms the inversion of configuration at the glycosidic linkage. Binding experiments show that the ADP-ribosylation has limited effect on nucleotide binding affinity, on ribosome binding, and on association with exotoxin A. These results provide insight to the inhibitory mechanism and suggest that inhibition may be caused by erroneous interaction of the translation factor with the codon-anticodon area in the P-site of the ribosome.
- Macromolecular Crystallography, Department of Molecular Biology, University of Aarhus, Gustav Wieds vej 10C, DK8000 Aarhus, Denmark.
Organizational Affiliation: