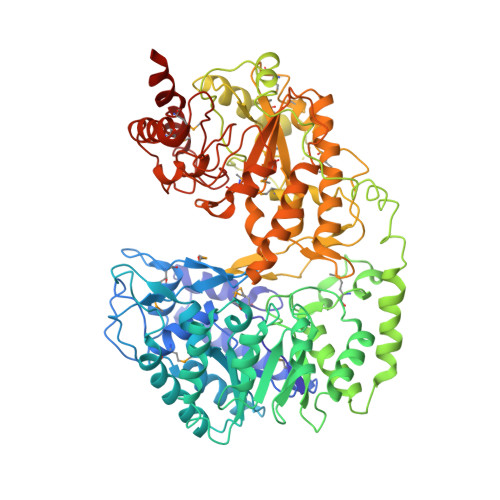
Crystal structures of cobalamin-independent methionine synthase complexed with zinc, homocysteine, and methyltetrahydrofolate
Ferrer, J.-L., Ravanel, S., Robert, M., Dumas, R.(2004) J Biological Chem 279: 44235-44238
- PubMed: 15326182
- DOI: https://doi.org/10.1074/jbc.C400325200
- Primary Citation Related Structures:
1U1H, 1U1J, 1U1U, 1U22 - PubMed Abstract:
Cobalamin-independent methionine synthase (MetE) catalyzes the synthesis of methionine by a direct transfer of the methyl group of N5-methyltetrahydrofolate (CH3-H2PteGlun) to the sulfur atom of homocysteine (Hcy). We report here the first crystal structure of this metalloenzyme under different forms, free or complexed with the Hcy and folate substrates. The Arabidopsis thaliana MetE (AtMetE) crystals reveal a monomeric structure built by two (betaalpha)8 barrels making a deep groove at their interface. The active site is located at the surface of the C-terminal domain, facing the large interdomain cleft. Inside the active site, His647, Cys649, and Cys733 are involved in zinc coordination, whereas Asp605, Ile437, and Ser439 interact with Hcy. Opposite the zinc/Hcy binding site, a cationic loop (residues 507-529) belonging to the C-terminal domain anchors the first glutamyl residue of CH3-H4PteGlu5. The pterin moiety of CH3-H4PteGlu5 is stacked with Trp567, enabling the N5-methyl group to protrude in the direction of the zinc atom. These data suggest a structural role of the N-terminal domain of AtMetE in the stabilization of loop 507-529 and in the interaction with the poly-glutamate chain of CH3-H4PteGlun. Comparison of AtMetE structures reveals that the addition of Hcy does not lead to a direct coordination of the sulfur atom with zinc but to a reorganization of the zinc binding site with a stronger coordination to Cys649, Cys733, and a water molecule.
- Laboratoire de Cristallogenèse et Cristallographie des Protéines, Institut de Biologie Structurale J.-P. Ebel, 41 rue Jules Horowitz, 38027 Grenoble 1, France. jean-luc.ferrer@ibs.fr
Organizational Affiliation: