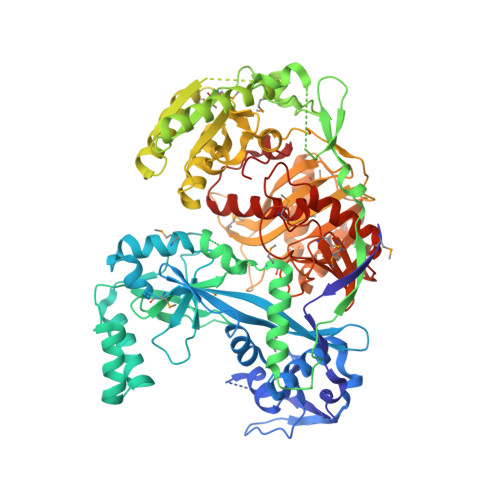
Crystal structure of Argonaute and its implications for RISC slicer activity.
Song, J.J., Smith, S.K., Hannon, G.J., Joshua-Tor, L.(2004) Science 305: 1434-1437
- PubMed: 15284453 Search on PubMed
- DOI: https://doi.org/10.1126/science.1102514
- Primary Citation Related Structures:
1U04 - PubMed Abstract:
Argonaute proteins and small interfering RNAs (siRNAs) are the known signature components of the RNA interference effector complex RNA-induced silencing complex (RISC). However, the identity of "Slicer," the enzyme that cleaves the messenger RNA (mRNA) as directed by the siRNA, has not been resolved. Here, we report the crystal structure of the Argonaute protein from Pyrococcus furiosus at 2.25 angstrom resolution. The structure reveals a crescent-shaped base made up of the amino-terminal, middle, and PIWI domains. The Piwi Argonaute Zwille (PAZ) domain is held above the base by a "stalk"-like region. The PIWI domain (named for the protein piwi) is similar to ribonuclease H, with a conserved active site aspartate-aspartate-glutamate motif, strongly implicating Argonaute as "Slicer." The architecture of the molecule and the placement of the PAZ and PIWI domains define a groove for substrate binding and suggest a mechanism for siRNA-guided mRNA cleavage.
- Watson School of Biological Sciences, 1 Bungtown Road, Cold Spring Harbor, NY 11724, USA.
Organizational Affiliation: