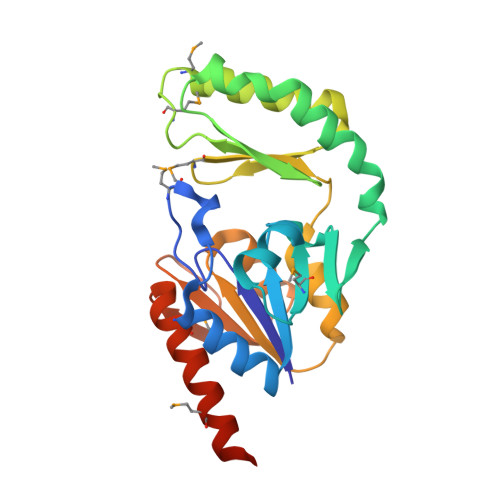
Crystal structure of trehalose-6-phosphate phosphatase-related protein: biochemical and biological implications.
Rao, K.N., Kumaran, D., Seetharaman, J., Bonanno, J.B., Burley, S.K., Swaminathan, S.(2006) Protein Sci 15: 1735-1744
- PubMed: 16815921 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062096606
- Primary Citation Related Structures:
1U02 - PubMed Abstract:
We report here the crystal structure of a trehalose-6-phosphate phosphatase-related protein (T6PP) from Thermoplasma acidophilum, TA1209, determined by the dual-wavelength anomalous diffraction (DAD) method. T6PP is a member of the haloacid dehalogenase (HAD) superfamily with significant sequence homology with trehalose-6-phosphate phosphatase, phosphoserine phosphatase, P-type ATPases and other members of the family. T6PP possesses a core domain of known alpha/beta-hydrolase fold, characteristic of the HAD family, and a cap domain, with a tertiary fold consisting of a four-stranded beta-sheet with two alpha-helices on one side of the sheet. An active-site magnesium ion and a glycerol molecule bound at the interface between the two domains provide insight into the mode of substrate binding by T6PP. A trehalose-6-phosphate molecule modeled into a cage formed by the two domains makes favorable interactions with the protein molecule. We have confirmed that T6PP is a trehalose phosphatase from amino acid sequence, three-dimensional structure, and biochemical assays.
- Biology Department, Brookhaven National Laboratory, Upton, NY 11973, USA.
Organizational Affiliation: