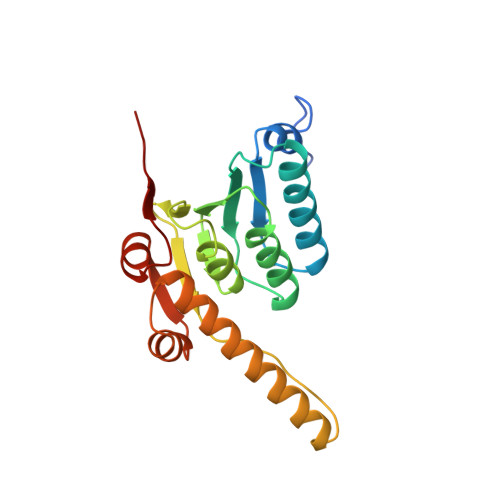
The structure of ClpP at 2.3 A resolution suggests a model for ATP-dependent proteolysis.
Wang, J., Hartling, J.A., Flanagan, J.M.(1997) Cell 91: 447-456
- PubMed: 9390554 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)80431-6
- Primary Citation Related Structures:
1TYF - PubMed Abstract:
We have determined the crystal structure of the proteolytic component of the caseinolytic Clp protease (ClpP) from E. coli at 2.3 A resolution using an ab initio phasing procedure that exploits the internal 14-fold symmetry of the oligomer. The structure of a ClpP monomer has a distinct fold that defines a fifth structural family of serine proteases but a conserved catalytic apparatus. The active protease resembles a hollow, solid-walled cylinder composed of two 7-fold symmetric rings stacked back-to-back. Its 14 proteolytic active sites are located within a central, roughly spherical chamber approximately 51 A in diameter. Access to the proteolytic chamber is controlled by two axial pores, each having a minimum diameter of approximately 10 A. From the structural features of ClpP, we suggest a model for its action in degrading proteins.
- Biology Department, Brookhaven National Laboratory, Upton, New York 11973-5000, USA.
Organizational Affiliation: