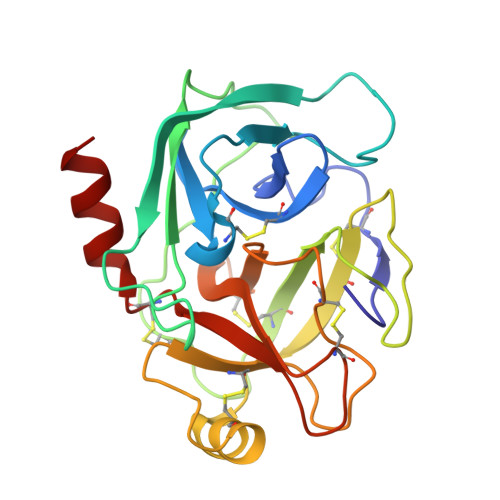
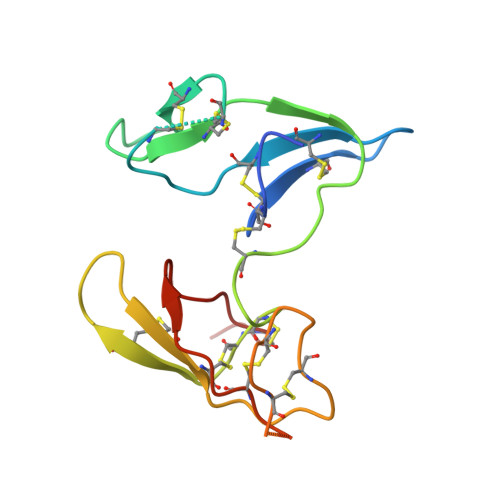
Crystal structure of the Bowman-Birk inhibitor from barley seeds in ternary complex with porcine trypsin
Park, E.Y., Kim, J.A., Kim, H.W., Kim, Y.S., Song, H.K.(2004) J Mol Biology 343: 173-186
- PubMed: 15381428 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.08.027
- Primary Citation Related Structures:
1TX6 - PubMed Abstract:
The structure and function of Bowman-Birk inhibitors (BBIs) from dicotyledonous plants such as soybean have been studied extensively. In contrast, relatively little is known about the BBIs from monocotyledonous plants such as barley, which differ from dicot BBIs in size and tertiary structure. The BBI from barley seeds (BBBI) consists of 125 amino acid residues with two separate inhibitory loops. Previously we determined the high-resolution structure of a 16 kDa BBBI in the free state. The BBBI folds into two compact domains (N and C domain) with tertiary structures that are similar to that of the 8 kDa BBI from dicots. Here we report the structure of a 1:2 complex between BBBI and porcine pancreatic trypsin (PPT) at 2.2 A resolution. This structure confirms that several regions, including the inhibitory loops in the free BBBI structure, show exceptionally low temperature factors and a distorted conformation due to crystalline packing in the lattice. Extensive analysis of the interaction between BBBI and trypsin, and comparison with other known canonical inhibitor-protease complexes, reveals that the mode of interaction between BBBI and PPT is similar to that of known serine protease inhibitors, as expected; however, several unique features are also identified in the primary binding sites near the inhibitory loops as well as in additional binding sites. The carboxy-terminal tail of the inhibitor extends into the interface between the two trypsin molecules and interacts with both of them simultaneously. The longest distance between the two P1 residues (Arg17 and Arg76) in the complex structure is approximately 34 A, which is shorter than in the free inhibitor, but it is still possible for BBBI to bind and inhibit two trypsin molecules simultaneously and independently.
- Research Institute, National Cancer Center, 809 Madu-dong, Ilsan-gu, Goyang-si, Gyeonggi-do 411-764, South Korea.
Organizational Affiliation: