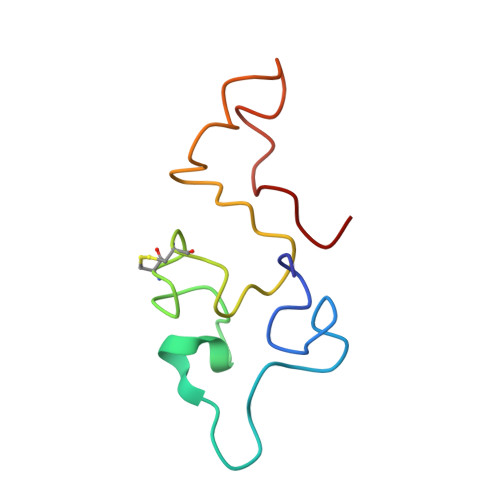
Structure of the equine infectious anemia virus Tat protein.
Willbold, D., Rosin-Arbesfeld, R., Sticht, H., Frank, R., Rosch, P.(1994) Science 264: 1584-1587
- PubMed: 7515512 Search on PubMed
- DOI: https://doi.org/10.1126/science.7515512
- Primary Citation Related Structures:
1TVT - PubMed Abstract:
Trans-activator (Tat) proteins regulate the transcription of lentiviral DNA in the host cell genome. These RNA binding proteins participate in the life cycle of all known lentiviruses, such as the human immunodeficiency viruses (HIV) or the equine infectious anemia virus (EIAV). The consensus RNA binding motifs [the trans-activation responsive element (TAR)] of HIV-1 as well as EIAV Tat proteins are well characterized. The structure of the 75-amino acid EIAV Tat protein in solution was determined by two- and three-dimensional nuclear magnetic resonance methods and molecular dynamics calculations. The protein structure exhibits a well-defined hydrophobic core of 15 amino acids that serves as a scaffold for two flexible domains corresponding to the NH2- and COOH-terminal regions. The core region is a strictly conserved sequence region among the known Tat proteins. The structural data can be used to explain several of the observed features of Tat proteins.
- Lehrstuhl für Biopolymere, Universität Bayreuth, Germany.
Organizational Affiliation: