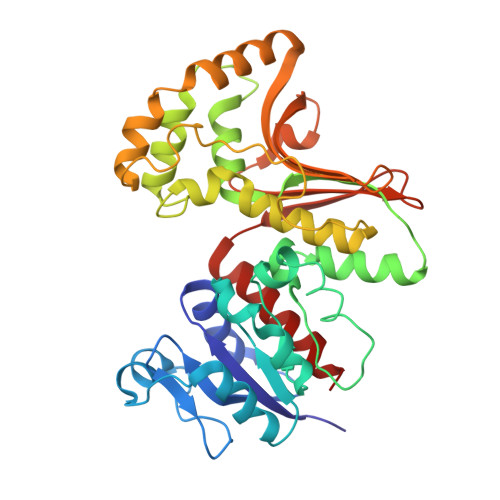
New phenolic inhibitors of yeast homoserine dehydrogenase
Ejim, L., Mirza, I.A., Capone, C., Nazi, I., Jenkins, S., Chee, G.L., Berghuis, A.M., Wright, G.D.(2004) Bioorg Med Chem 12: 3825-3830
- PubMed: 15210149
- DOI: https://doi.org/10.1016/j.bmc.2004.05.009
- Primary Citation of Related Structures:
1TVE - PubMed Abstract:
A relatively unexploited potential target for antimicrobial agents is the biosynthesis of essential amino acids. Homoserine dehydrogenase, which reduces aspartate semi-aldehyde to homoserine in a NAD(P)H-dependent reaction, is one such target that is required for the biosynthesis of Met, Thr, and Ile from Asp. We report a small molecule screen of yeast homoserine dehydrogenase that has identified a new class of phenolic inhibitors of this class of enzyme. X-ray crystal structural analysis of one of the inhibitors in complex with homoserine dehydrogenase reveals that these molecules bind in the amino acid binding region of the active site and that the phenolic hydroxyl group interacts specifically with the backbone amide of Gly175. These results provide the first nonamino acid inhibitors of this class of enzyme and have the potential to be exploited as leads in antifungal compound design.
- Antimicrobial Research Centre, Department of Biochemistry, McMaster University, 1200 Main Street West, Hamilton, ON, Canada L8N 3Z5.
Organizational Affiliation: