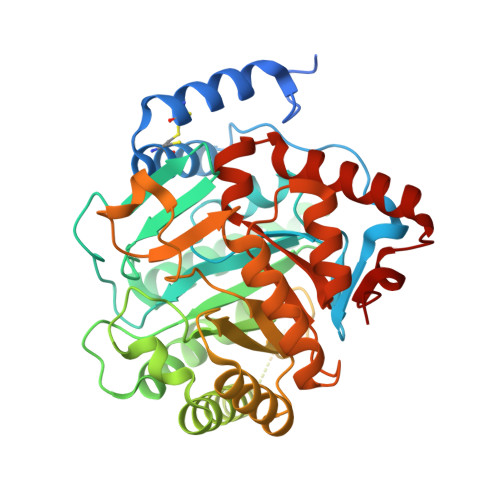
Structure of Plasmodium falciparum dihydroorotate dehydrogenase with a bound inhibitor.
Hurt, D.E., Widom, J., Clardy, J.(2006) Acta Crystallogr D Biol Crystallogr 62: 312-323
- PubMed: 16510978 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905042642
- Primary Citation Related Structures:
1TV5 - PubMed Abstract:
Membrane-associated dihydroorotate dehydrogenase (DHODH) is an antimalarial therapeutic target without an effective inhibitor. Studies on human DHODH (HsDHODH) led to a structural mechanistic model in which respiratory quinones bind in a tunnel formed by the highly variable N-terminus that leads to the flavin mononucleotide-binding site. The therapeutic agents leflunomide (Arava) and brequinar sodium inhibit HsDHODH by binding in this tunnel. Plasmodium falciparum DHODH (PfDHODH) and HsDHODH have markedly different sensitivities to the two drugs. To understand the structural basis of this differential sensitivity and begin a structure-based drug-design cycle for PfDHODH inhibitors, the three-dimensional structure (2.4 Angstroms, R = 20.1%) of PfDHODH bound to the active metabolite of leflunomide was determined by X-ray crystallography. Comparison of the structures of HsDHODH and PfDHODH reveals a completely different binding mode for the same inhibitor in these two catalytically identical enzymes and explains the previously observed species-specific preferential binding. Because no effective inhibitors have been described for PfDHODH, this structure provides critical insight for the design of potential antimalarials.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, NY 14850, USA.
Organizational Affiliation: