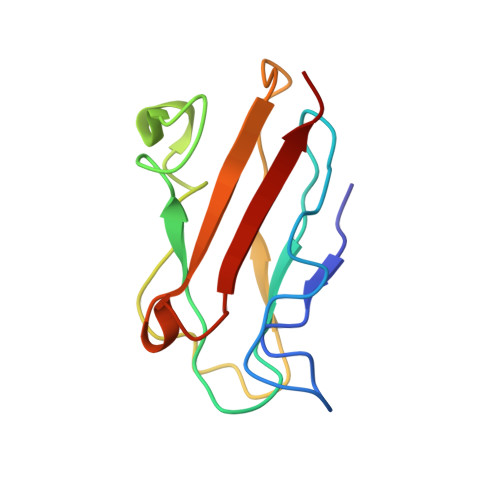
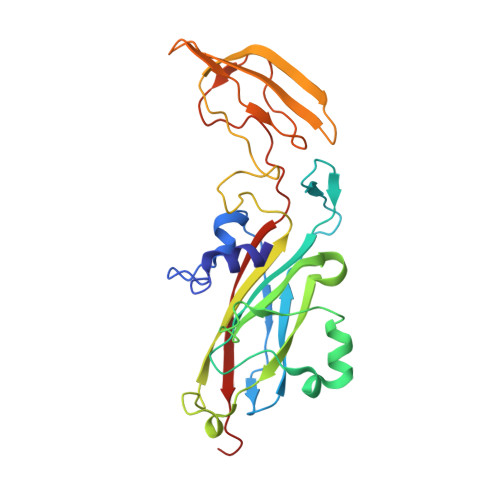
Structure of the complex between plastocyanin and cytochrome f from the cyanobacterium Nostoc sp. PCC 7119 as determined by paramagnetic NMR. The balance between electrostatic and hydrophobic interactions within the transient complex determines the relative orientation of the two proteins.
Diaz-Moreno, I., Diaz-Quintana, A., De la Rosa, M.A., Ubbink, M.(2005) J Biological Chem 280: 18908-18915
- PubMed: 15705583
- DOI: https://doi.org/10.1074/jbc.M413298200
- Primary Citation of Related Structures:
1TU2 - PubMed Abstract:
The complex between cytochrome f and plastocyanin from the cyanobacterium Nostoc has been characterized by NMR spectroscopy. The binding constant is 16 mM(-1), and the lifetime of the complex is much less than 10 ms. Intermolecular pseudo-contact shifts observed for the plastocyanin amide nuclei, caused by the heme iron, as well as the chemical-shift perturbation data were used as the sole experimental restraints to determine the orientation of plastocyanin relative to cytochrome f with a precision of 1.3 angstroms. The data show that the hydrophobic patch surrounding tyrosine 1 in cytochrome f docks the hydrophobic patch of plastocyanin. Charge complementarities are found between the rims of the respective recognition sites of cytochrome f and plastocyanin. Significant differences in the relative orientation of both proteins are found between this complex and those previously reported for plants and Phormidium, indicating that electrostatic and hydrophobic interactions are balanced differently in these complexes.
- Instituto de Bioquímica Vegetal y Fotosíntesis, Universidad de Sevilla y Consejo Superior de Investigaciones Científicas, Avda. Américo Vespucio 49, 41092 Sevilla, Spain.
Organizational Affiliation: