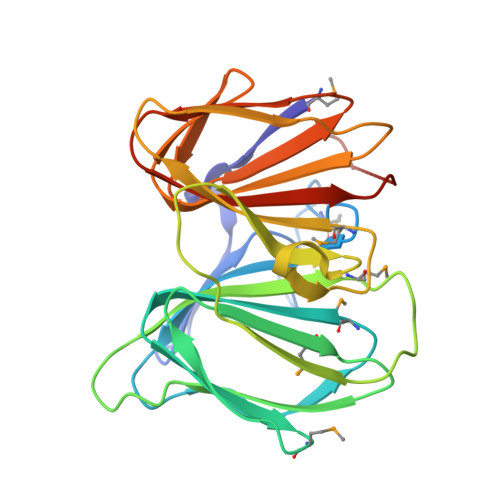
Structural and biochemical analysis reveal pirins to possess quercetinase activity.
Adams, M., Jia, Z.(2005) J Biological Chem 280: 28675-28682
- PubMed: 15951572
- DOI: https://doi.org/10.1074/jbc.M501034200
- Primary Citation Related Structures:
1TQ5 - PubMed Abstract:
Pirin is a recently identified eukaryotic protein implicated in transcriptional activation and apoptosis. Homologues of Pirin are highly conserved in both prokaryotes and eukaryotes, but their function remains poorly understood. We present here the crystal structure of the yhhW gene product, a putative Pirin homologue, from Escherichia coli and confirm its structural similarity to Pirin. The YhhW protein displays a bicupin fold with a single N-terminal metal coordination site. Molecular surface comparisons of YhhW and Pirin with structurally similar proteins suggested quercetin as a potential ligand. We demonstrate that both bacterial and human Pirins have quercetinase activity, which is inhibited by the addition of typical inhibitors of the quercetin 2,3-dioxygenase reaction. We also demonstrate the release of carbon monoxide as a reaction product. This is the first report of enzymatic activity for any member of the Pirin family and may be an important connection to their roles in transcriptional regulation.
- Department of Biochemistry, Queen's University, Kingston, Ontario K7L 3N6, Canada.
Organizational Affiliation: