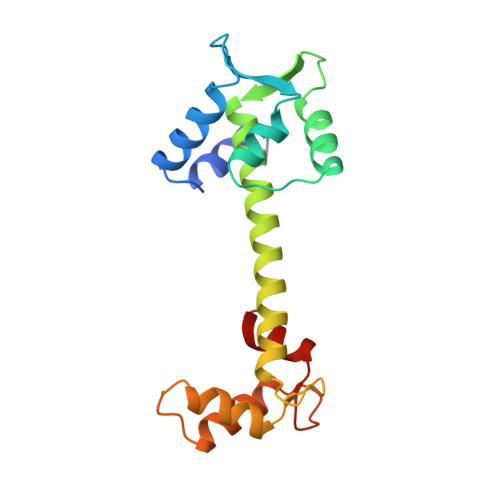
Structure of chicken skeletal muscle troponin C at 1.78 A resolution.
Satyshur, K.A., Pyzalska, D., Greaser, M., Rao, S.T., Sundaralingam, M.(1994) Acta Crystallogr D Biol Crystallogr 50: 40-49
- PubMed: 15299475 Search on PubMed
- DOI: https://doi.org/10.1107/S090744499300798X
- Primary Citation Related Structures:
1TOP - PubMed Abstract:
The structure of chicken skeletal muscle troponin C (TnC) has been refined to an R value of 0.168, using 14 788 reflections, in the resolution range 8.0-1.78 A. Our earlier 2 A resolution structure [Satyshur, Rao, Pyzalska, Drendel, Greaser & Sundaralingam (1988). J. Biol. Chem. 263, 1628-1647] served as the starting model. The refined model includes atoms for all protein residues (1-162), 2 Ca(2+) ions, 169 water molecules and one sulfate ion. The high-resolution refinement shows more clearly the details of the protein and water structure. The side chains Glu63, Cysl01, Arg123, Aspl40 and Asp152 adopt two discretely ordered conformations. The long central helix is only slightly curved/bent (7.9 degrees ) and all the central helix NH.O=C hydrogen bonds are intact. Seven of the nine carbonyl O atoms of the mid segment of this helix, including the D/E linker region, are hydrogen bonded to water molecules which weakens the helix hydrogen bonds. In contrast, in each of the protected upper and lower thirds of the long central helix, only two carbonyl O atoms are hydrogen bonded to water molecules. The hydrogen-bonding patterns displayed by some of the carbonyl O atoms of NT and A helices of the N-terminal domain and the F and H helices of the C-terminal domain, which are on the exposed surface of the protein, are similar. The B helix of the calcium-free site I is kinked, with the local helix axes at either end making an angle of 39 degrees, by two inserted water molecules between N-H and O=C groups, breaking the adjacent helix hydrogen bonds. A sulfate ion from the crystallization buffer is also trapped in the B helix between the guanidinium group of Arg47 and these two inserted water molecules. The C helix of site II is devoid of similar hydration and is probably responsible for the different interhelical angles A/B at site I (134 degrees ) and C/D at site II (149 degrees ). Extensive interhelix hydrogen bonds occur between the side chains of the C and D helices of the 'apo' site II: Gln51-Asp89, Asn52-Asp89, Glu57-Gln85, Glu57-Glu88 and Glu64-Arg84, which apparently are disrupted upon Ca uptake and the resulting rearrangement of the helices expose the side chains, lining the palm of the N-(and C-) terminal domains, for interaction with specific peptide fragment of troponin I (Tnl) during muscle contraction. The dominant crystal packing motif involves a head-to-tail interaction between the N-terminal domain A helix of one molecule and the palm of the C-terminal domain of the 3(2)-related molecule, in a manner similar to that which can be expected for the TnC-TnI complex. Similar interactions may also be responsible for the dimerization of TnC at low pH.
- Crystallography Laboratory, Department of Biochemistry and College of Agriculture and Life Sciences, University of Wisconsin, Madison 53706, USA.
Organizational Affiliation: