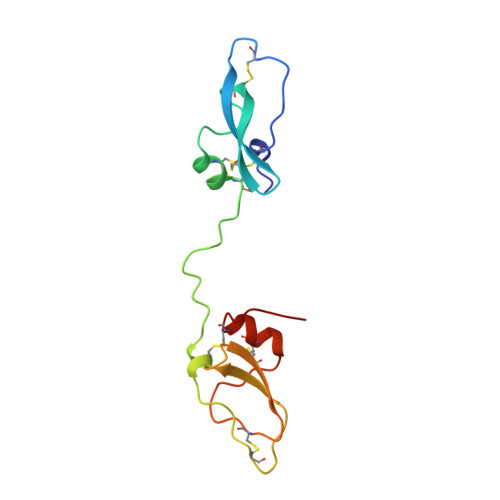
The ornithodorin-thrombin crystal structure, a key to the TAP enigma?
van de Locht, A., Stubbs, M.T., Bode, W., Friedrich, T., Bollschweiler, C., Hoffken, W., Huber, R.(1996) EMBO J 15: 6011-6017
- PubMed: 8947023 Search on PubMedSearch on PubMed Central
- Primary Citation Related Structures:
1TOC - PubMed Abstract:
Ornithodorin, isolated from the blood sucking soft tick Ornithodoros moubata, is a potent (Ki = 10(-12) M) and highly selective thrombin inhibitor. Internal sequence homology indicates a two domain protein. Each domain resembles the Kunitz inhibitor basic pancreatic trypsin inhibitor (BPTI) and also the tick anticoagulant peptide (TAP) isolated from the same organism. The 3.1 A crystal structure of the ornithodorin-thrombin complex confirms that both domains of ornithodorin exhibit a distorted BPTI-like fold. The N-terminal portion and the C-terminal helix of each domain are structurally very similar to BPTI, whereas the regions corresponding to the binding loop of BPTI adopt different conformations. Neither of the two 'reactive site loops' of ornithodorin contacts the protease in the ornithodorin-thrombin complex. Instead, the N-terminal residues of ornithodorin bind to the active site of thrombin, reminiscent of the thrombin-hirudin interaction. The C-terminal domain binds at the fibrinogen recognition exosite. Molecular recognition of its target protease by this double-headed Kunitz-type inhibitor diverges considerably from other members of this intensely studied superfamily. The complex structure provides a model to explain the perplexing results of mutagenesis studies on the TAP-factor Xa interaction.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Martinsreid, Germany.
Organizational Affiliation: