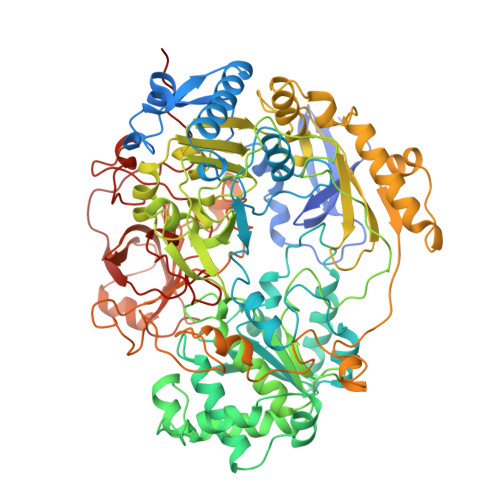
Crystal structure of oxidized trimethylamine N-oxide reductase from Shewanella massilia at 2.5 A resolution.
Czjzek, M., Dos Santos, J.P., Pommier, J., Giordano, G., Mejean, V., Haser, R.(1998) J Mol Biology 284: 435-447
- PubMed: 9813128
- DOI: https://doi.org/10.1006/jmbi.1998.2156
- Primary Citation of Related Structures:
1TMO - PubMed Abstract:
The periplasmic trimethylamine N-oxide (TMAO) reductase from the marine bacteria Shewanella massilia is involved in a respiratory chain, having trimethylamine N-oxide as terminal electron acceptor. This molybdoenzyme belongs to the dimethyl sulfoxide (DMSO) reductase family, but has a different substrate specificity than its homologous enzyme. While the DMSO reductases reduce a broad spectra of organic S-oxide and N-oxide compounds, TMAO reductase from Shewanella massilia reduces only TMAO as the natural compound. The crystal structure was solved by molecular replacement with the coordinates of the DMSO reductase from Rhodobacter sphaeroides. The overall fold of the protein structure is essentially the same as the DMSO reductase structures, organized into four domains. The molybdenum coordination sphere is closest to that described in the DMSO reductase of Rhodobacter capsulatus. The structural differences found in the protein environment of the active site could be related to the differences in substrate specificity of these enzymes. In close vicinity of the molybdenum ion a tyrosine residue is missing in the TMAO reductase, leaving a greater space accessible to the solvent. This tyrosine residue has contacts to the oxo groups in the DMSO reductase structures. The arrangement and number of charged residues lining the inner surface of the funnel-like entrance to the active site, is different in the TMAO reductase than in the DMSO reductases from Rhodobacter species. Furthermore a surface loop at the top of the active-site funnel, for which no density was present in the DMSO reductase structures, is well defined in the oxidized form of the TMAO reductase structure, and is located on the border of the funnel-like entrance of the active center.
- Laboratoire d'Architecture et Fonction de Macromolécules Biologiques, AFMB-CNRS Marseille, IBSM, 31 chemin Joseph Aiguier, Marseille Cedex 20, 13402, France. czjzek@afmb.cnrs-mrs.fr
Organizational Affiliation: