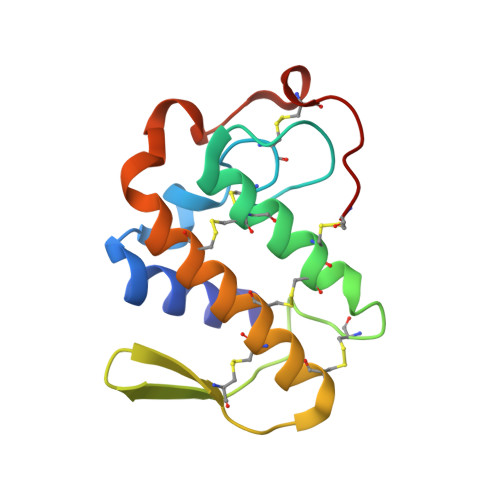
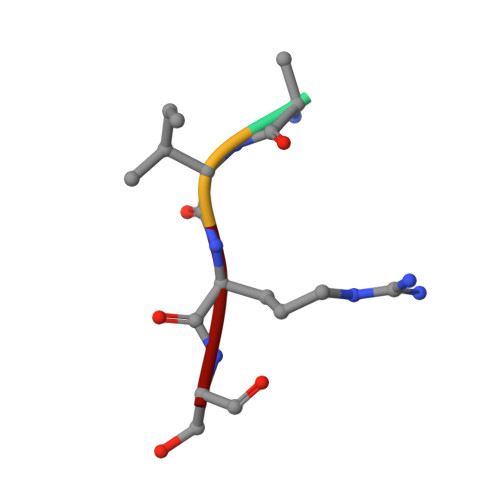
Crystal structure of russells viper phospholipase A2 with a specifically designed tetrapeptide Ala-Ile-Arg-Ser at 1.1 A resolution
Singh, N., Bilgrami, S., Somvanshi, R.K., Sharma, S., Dey, S., Perbandt, M., Betzel, C., Kaur, P., Singh, T.P.To be published.