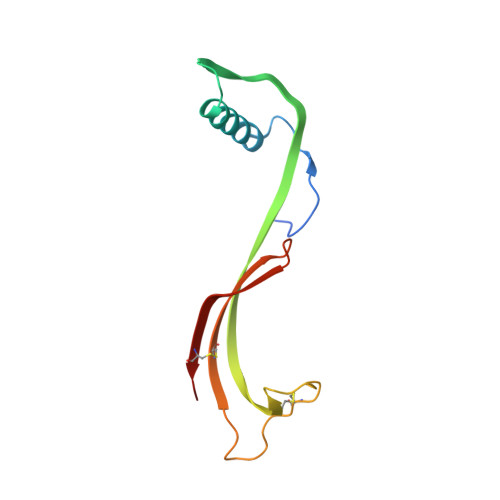
3D domain-swapped human cystatin C with amyloidlike intermolecular beta-sheets.
Janowski, R., Kozak, M., Abrahamson, M., Grubb, A., Jaskolski, M.(2005) Proteins 61: 570-578
- PubMed: 16170782 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20633
- Primary Citation Related Structures:
1TIJ - PubMed Abstract:
Oligomerization of human cystatin C (HCC) leads to amyloid deposits in brain arteries, and this process is greatly accelerated with a naturally occurring L68Q variant. The crystal structures of N-truncated and full-length HCC (cubic form) showed dimer formation via three-dimensional (3D) domain swapping, and this observation has led to the suggestion that an analogous domain-swapping mechanism, but propagated in an open-ended fashion, could be the basis of HCC fibril formation. Here we report that full-length HCC, when crystallized in a new, tetragonal form, dimerizes by swapping the same secondary structure elements but with a very different overall structure generated by the flexibility of the hinge linking the moveable elements. The beta-strands of the beta-cores of the two folding units of the present dimer are roughly parallel, while they formed an angle of about 100 degrees in the previous two structures. The dimers pack around a crystallographic dyad by extending their molecular beta-sheets in an intermolecular context. At the other edge of the molecular beta-sheet, side-chain-side-chain hydrogen bonds propagate the beta-structure in the same direction. In consequence, a supramolecular crystal structure is generated, with all the beta-strands of the domain-swapped dimers being perpendicular to one crystallographic direction. This observation is relevant to amyloid aggregation of HCC, as X-ray diffraction studies of amyloid fibrils show them to have ordered, repeating structure, consistent with the so-called cross-beta structure, in which extended polypeptide chains are perpendicular to the fiber axis and form infinite beta-sheets that are parallel to this axis.
- Department of Crystallography, Faculty of Chemistry, A. Mickiewicz University, Poznan, Poland.
Organizational Affiliation: