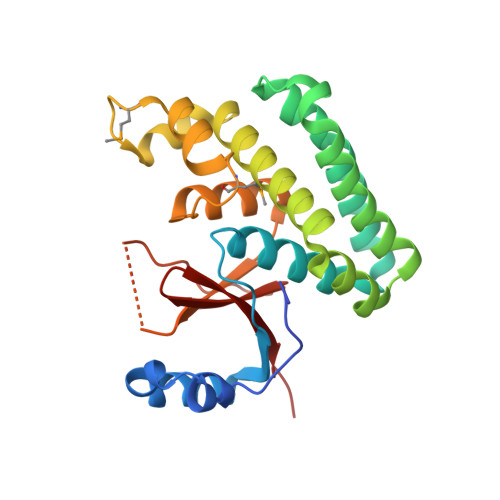
Crystal structure of human otubain 2.
Nanao, M.H., Tcherniuk, S.O., Chroboczek, J., Dideberg, O., Dessen, A., Balakirev, M.Y.(2004) EMBO Rep 5: 783-788
- PubMed: 15258613 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.embor.7400201
- Primary Citation Related Structures:
1TFF - PubMed Abstract:
Ubiquitylation, the modification of cellular proteins by the covalent attachment of ubiquitin, is critical for diverse biological processes including cell cycle progression, signal transduction and stress response. This process can be reversed and regulated by a group of proteases called deubiquitylating enzymes (DUBs). Otubains are a recently identified family of DUBs that belong to the ovarian tumour (OTU) superfamily of proteins. Here, we report the first crystal structure of an OTU superfamily protein, otubain 2, at 2.1 A resolution and propose a model for otubain-ubiquitin binding on the basis of other DUB structures. Although otubain 2 is a member of the cysteine protease superfamily of folds, its crystal structure shows a novel fold for DUBs. Moreover, the active-site cleft is sterically occluded by a novel loop conformation resulting in an oxyanion hole, which consists uniquely of backbone amides, rather than the composite backbone/side-chain substructures seen in other DUBs and cysteine proteases. Furthermore, the residues that orient and stabilize the active-site histidine of otubain 2 are different from other cysteine proteases. This reorganization of the active-site topology provides a possible explanation for the low turnover and substrate specificity of the otubains.
- Laboratoire de Cristallographie Macromoléculaire, Institut de Biologie Structurale JP Ebel (CEA/CNRS/UJF), 41 rue Jules Horowitz, 38027 Grenoble, France.
Organizational Affiliation: