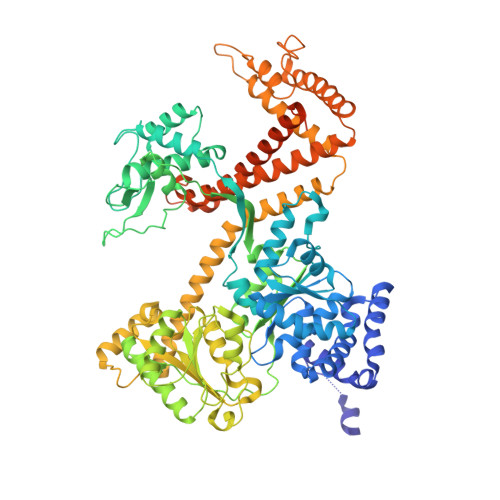
A large conformational change of the translocation ATPase SecA.
Osborne, A.R., Clemons Jr., W.M., Rapoport, T.A.(2004) Proc Natl Acad Sci U S A 101: 10937-10942
- PubMed: 15256599 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0401742101
- Primary Citation Related Structures:
1TF2, 1TF5 - PubMed Abstract:
The ATPase SecA mediates the posttranslational translocation of a wide range of polypeptide substrates through the SecY channel in the cytoplasmic membrane of bacteria. We have determined the crystal structure of a monomeric form of Bacillus subtilis SecA at a 2.2-A resolution. A comparison with the previously determined structures of SecA reveals a nucleotide-independent, large conformational change that opens a deep groove similar to that in other proteins that interact with diverse polypeptides. We propose that the open form of SecA represents an activated state.
- Department of Cell Biology, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: