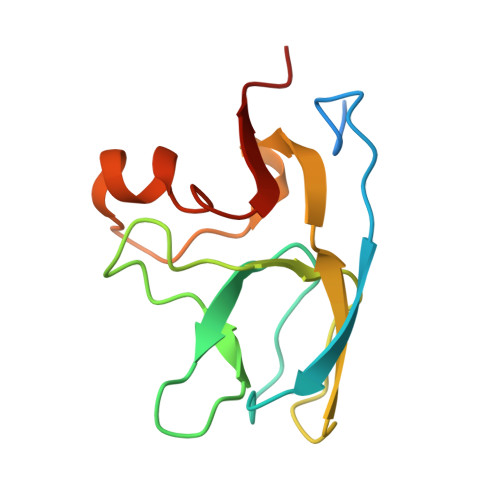
Kinetic Stability and Crystal Structure of the Viral Capsid Protein SHP.
Forrer, P., Chang, C., Ott, D., Wlodawer, A., Plueckthun, A.(2004) J Mol Biology 344: 179-193
- PubMed: 15504410
- DOI: https://doi.org/10.1016/j.jmb.2004.09.030
- Primary Citation of Related Structures:
1TD0, 1TD3, 1TD4 - PubMed Abstract:
SHP, the capsid-stabilizing protein of lambdoid phage 21, is highly resistant against denaturant-induced unfolding. We demonstrate that this high functional stability of SHP is due to a high kinetic stability with a half-life for unfolding of 25 days at zero denaturant, while the thermodynamic stability is not unusually high. Unfolding experiments demonstrated that the trimeric state (also observed in crystals and present on the phage capsid) of SHP is kinetically stable in solution, while the monomer intermediate unfolds very rapidly. We also determined the crystal structure of trimeric SHP at 1.5A resolution, which was compared to that of its functional homolog gpD. This explains how a tight network of H-bonds rigidifies crucial interpenetrating residues, leading to the observed extremely slow trimer dissociation or denaturation. Taken as a whole, our results provide molecular-level insights into natural strategies to achieve kinetic stability by taking advantage of protein oligomerization. Kinetic stability may be especially needed in phage capsids to allow survival in harsh environments.
- Biochemisches Institut, Universität Zürich, Winterthurerstrasse 190, CH-8057 Zürich, Switzerland.
Organizational Affiliation: