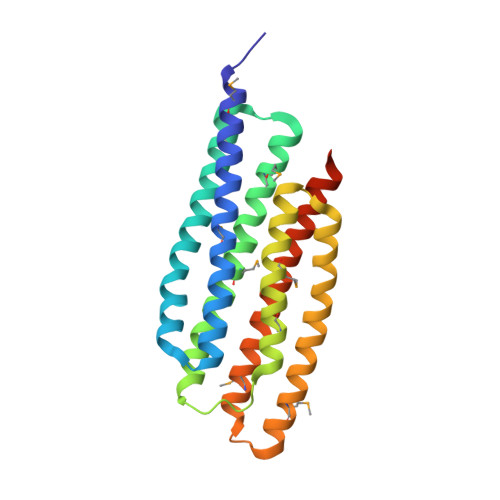
Crystal structure of the "PhoU-like" phosphate uptake regulator from Aquifex aeolicus.
Oganesyan, V., Oganesyan, N., Adams, P.D., Jancarik, J., Yokota, H.A., Kim, R., Kim, S.H.(2005) J Bacteriol 187: 4238-4244
- PubMed: 15937186
- DOI: https://doi.org/10.1128/JB.187.12.4238-4244.2005
- Primary Citation Related Structures:
1T72, 1T8B - PubMed Abstract:
The phoU gene of Aquifex aeolicus encodes a protein called PHOU_AQUAE with sequence similarity to the PhoU protein of Escherichia coli. Despite the fact that there is a large number of family members (more than 300) attributed to almost all known bacteria and despite PHOU_AQUAE's association with the regulation of genes for phosphate metabolism, the nature of its regulatory function is not well understood. Nearly one-half of these PhoU-like proteins, including both PHOU_AQUAE and the one from E. coli, form a subfamily with an apparent dimer structure of two PhoU domains on the basis of their amino acid sequence. The crystal structure of PHOU_AQUAE (a 221-amino-acid protein) reveals two similar coiled-coil PhoU domains, each forming a three-helix bundle. The structures of PHOU_AQUAE proteins from both a soluble fraction and refolded inclusion bodies (at resolutions of 2.8 and 3.2A, respectively) showed no significant differences. The folds of the PhoU domain and Bag domains (for a class of cofactors of the eukaryotic chaperone Hsp70 family) are similar. Accordingly, we propose that gene regulation by PhoU may occur by association of PHOU_AQUAE with the ATPase domain of the histidine kinase PhoR, promoting release of its substrate PhoB. Other proteins that share the PhoU domain fold include the coiled-coil domains of the STAT protein, the ribosome-recycling factor, and structural proteins like spectrin.
- Berkely Structural Genomics Center, Physical Biosciences Division, Lawrence Berkeley National Laboratory, 1 Cyclotron Rd., Berkeley, California 94720, USA.
Organizational Affiliation: