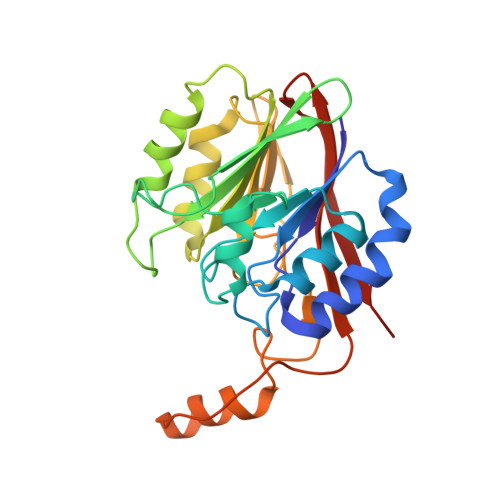
Structural and enzymatic characterization of DR1281: A calcineurin-like phosphoesterase from Deinococcus radiodurans.
Shin, D.H., Proudfoot, M., Lim, H.J., Choi, I.K., Yokota, H., Yakunin, A.F., Kim, R., Kim, S.H.(2008) Proteins 70: 1000-1009
- PubMed: 17847097 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21584
- Primary Citation Related Structures:
1T70 - PubMed Abstract:
We have determined the crystal structure of DR1281 from Deinococcus radiodurans. DR1281 is a protein of unknown function with over 170 homologs found in prokaryotes and eukaryotes. To elucidate the molecular function of DR1281, its crystal structure at 2.3 A resolution was determined and a series of biochemical screens for catalytic activity was performed. The crystal structure shows that DR1281 has two domains, a small alpha domain and a putative catalytic domain formed by a four-layered structure of two beta-sheets flanked by five alpha-helices on both sides. The small alpha domain interacts with other molecules in the asymmetric unit and contributes to the formation of oligomers. The structural comparison of the putative catalytic domain with known structures suggested its biochemical function to be a phosphatase, phosphodiesterase, nuclease, or nucleotidase. Structural analyses with its homologues also indicated that there is a dinuclear center at the interface of two domains formed by Asp8, Glu37, Asn38, Asn65, His148, His173, and His175. An absolute requirement of metal ions for activity has been proved by enzymatic assay with various divalent metal ions. A panel of general enzymatic assays of DR1281 revealed metal-dependent catalytic activity toward model substrates for phosphatases (p-nitrophenyl phosphate) and phosphodiesterases (bis-p-nitrophenyl phosphate). Subsequent secondary enzymatic screens with natural substrates demonstrated significant phosphatase activity toward phosphoenolpyruvate and phosphodiesterase activity toward 2',3'-cAMP. Thus, our structural and enzymatic studies have identified the biochemical function of DR1281 as a novel phosphatase/phosphodiesterase and disclosed key conserved residues involved in metal binding and catalytic activity.
- College of Pharmacy, Ewha Womans University, Seoul 120-750, Republic of Korea. dhshin55@ewha.ac.kr
Organizational Affiliation: