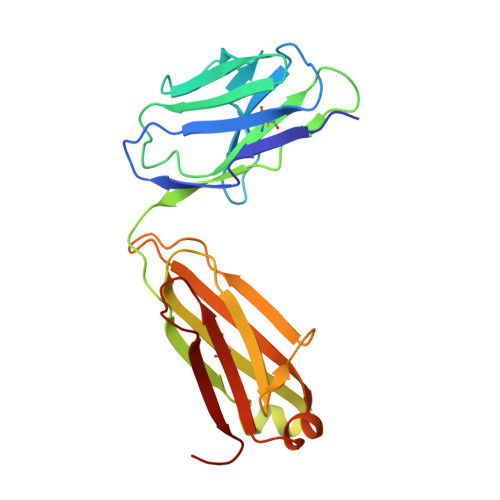
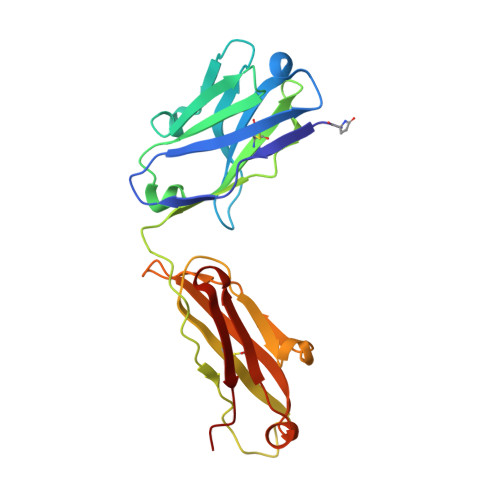
Three-dimensional structures of a humanized anti-IFN-gamma Fab (HuZAF) in two crystal forms.
Bourne, P.C., Terzyan, S.S., Cloud, G., Landolfi, N.F., Vasquez, M., Edmundson, A.B.(2004) Acta Crystallogr D Biol Crystallogr 60: 1761-1769
- PubMed: 15388922
- DOI: https://doi.org/10.1107/S0907444904018670
- Primary Citation of Related Structures:
1T04, 1T3F - PubMed Abstract:
Three-dimensional structures were determined for two crystal forms (orthorhombic P2(1)2(1)2(1) and monoclinic C2) of the Fab from the humanized version of a murine monoclonal antibody (AF2) that possesses binding and potent neutralizing activity against human interferon gamma (IFN-gamma). This humanized antibody (HuZAF; USAN name fontolizumab) is currently in phase II clinical trials for the treatment of Crohn's disease. HuZAF exhibits binding and IFN-gamma neutralizing capacities that closely approximate those of the original antibody. It is shown that HuZAF, whose VH domain was designed using a best-sequence-fit approach, is closer structurally to its mouse precursor than is a version whose VH was constructed using a human sequence with lower homology to the original mouse sequence. This work thus offers direct structural evidence in support of the best-sequence-fit approach and adds to previous results of biological and biochemical evaluations of distinctly engineered antibodies that also favored the use of a best-sequence-fit strategy. A second crystal type appeared during attempts to crystallize the Fab-IFN-gamma complex. The antibody-antigen complex that existed in solution dissociated in the crystallization mixture. A conformationally altered but unliganded HuZAF protein crystallized in a different space group (C2), with two Fab molecules in the asymmetric unit. In this crystal lattice, no space was available for accommodating the IFN-gamma antigen. Thus, there are currently three slightly different structures of the HuZAF Fab.
- Department of Veterinary Pathobiology, Oklahoma State University, Stillwater, OK 74078-2007, USA.
Organizational Affiliation: