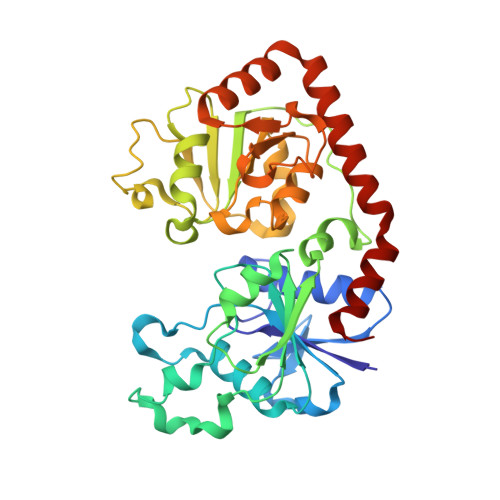
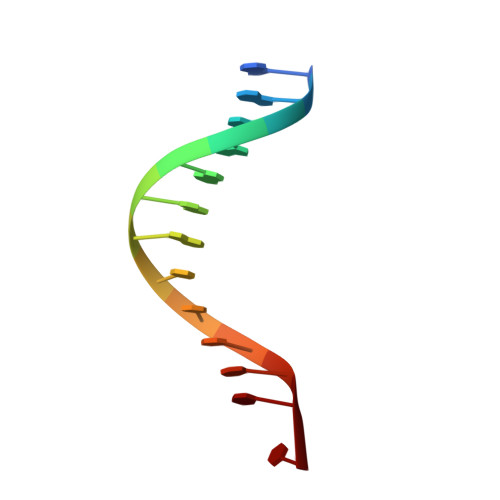
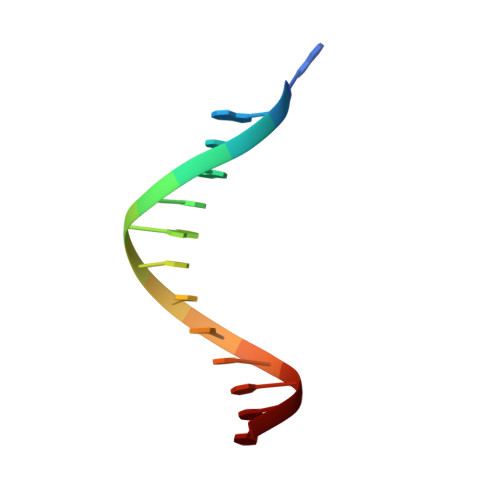
Structural evidence of a passive base flipping mechanism for {beta}-Glucosyltransferase
Lariviere, L., Morera, S.(2004) J Biological Chem 279: 34715-34720
- PubMed: 15178685 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M404394200
- Primary Citation Related Structures:
1SXP, 1SXQ - PubMed Abstract:
Beta-glucosyltransferase (BGT) is a DNA-modifying enzyme and a glycosyltransferase. This inverting enzyme transfers glucose from UDP-glucose to the 5-hydroxymethyl cytosine bases of T4 phage DNA. From previous structural analyses we showed that Asp-100 and Asn-70 were, respectively, the catalytic base and the key residue for specific DNA recognition (Larivière, L., Gueguen-Chaignon, V., and Moréra, S. (2003) J. Mol. Biol. 330, 1077-1086). Here, we supply biochemical evidence supporting their essential roles in catalysis. We have also shown previously that BGT uses a base-flipping mechanism to access 5-hydroxymethyl cytosine (Larivière, L., and Moréra, S. (2002) J. Mol. Biol. 324, 483-490). Whether it is an active or a passive process remains unclear, as is the case for all DNA cleaving and modifying enzymes. Here, we report two crystal structures: (i) BGT in complex with a 13-mer DNA containing an A:G mismatch and (ii) BGT in a ternary complex with UDP and an oligonucleotide containing a single central G:C base pair. The binary structure reveals a specific complex with the flipped-out, mismatched adenine exposed to the active site. Unexpectedly, the other structure shows the non-productive binding of an intermediate flipped-out base. Our structural analysis provides clear evidence for a passive process.
- Laboratoire d'Enzymologie et Biochimie Structurales, Unité Propre de Recherche 9063 CNRS, Bātiment 34, 1 Avenue de la Terrasse, 91198 Gif-sur-Yvette, France.
Organizational Affiliation: