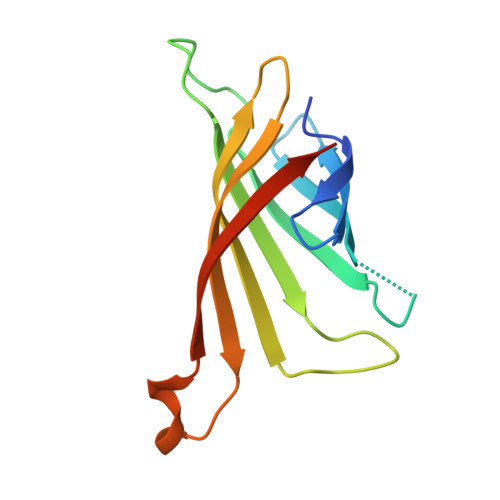
Structural studies of binding site tryptophan mutants in the high-affinity streptavidin-biotin complex.
Freitag, S., Le Trong, I., Chilkoti, A., Klumb, L.A., Stayton, P.S., Stenkamp, R.E.(1998) J Mol Biology 279: 211-221
- PubMed: 9636711 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.1735
- Primary Citation Related Structures:
1SWH, 1SWJ, 1SWK, 1SWL, 1SWN, 1SWO, 1SWP, 1SWQ, 1SWR - PubMed Abstract:
Previous thermodynamic and computational studies have pointed to the important energetic role of aromatic contacts in generating the exceptional binding free energy of streptavidin-biotin association. We report here the crystallographic characterization of single site tryptophan mutants in investigating structural consequences of alterations in these aromatic contacts. Four tryptophan residues, Trp79, Trp92, Trp108 and Trp120, play an important role in the hydrophobic binding contributions, which along with a hydrogen bonding network and a flexible binding loop give rise to tight ligand binding (Ka approximately 10(13) M-1). The crystal structures of ligand-free and biotin-bound mutants, W79F, W108F, W120F and W120A, in the resolution range from 1.9 to 2.3 A were determined. Nine data sets for these four different mutants were collected, and structural models were refined to R-values ranging from 0.15 to 0.20. The major question addressed here is how these mutations influence the streptavidin binding site and in particular how they affect the binding mode of biotin in the complex. The overall folding of streptavidin was not significantly altered in any of the tryptophan mutants. With one exception, only minor deviations in the unbound structures were observed. In one crystal form of unbound W79F, there is a coupled shift in the side-chains of Phe29 and Tyr43 toward the mutation site, although in a different crystal form these shifts are not observed. In the bound structures, the orientation of biotin in the binding pocket was not significantly altered in the mutant complex. Compared with the wild-type streptavidin-biotin complex, there were no additional crystallographic water molecules observed for any of the mutants in the binding pocket. These structural studies thus suggest that the thermodynamic alterations can be attributed to the local alterations in binding residue composition, rather than a rearrangement of binding site architectures.
- Department of Biological Structure, University of Washington, Seattle 98195-7420, USA.
Organizational Affiliation: