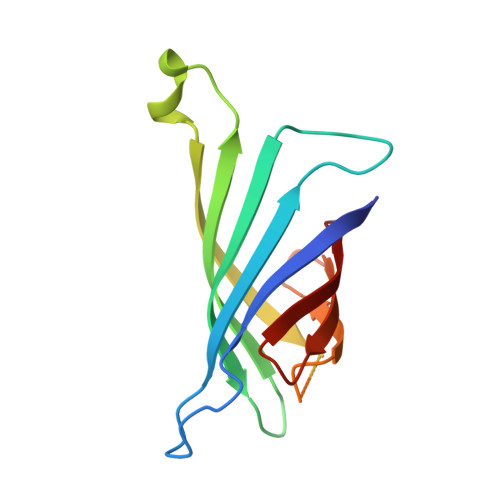
Thermodynamic and structural consequences of flexible loop deletion by circular permutation in the streptavidin-biotin system.
Chu, V., Freitag, S., Le Trong, I., Stenkamp, R.E., Stayton, P.S.(1998) Protein Sci 7: 848-859
- PubMed: 9568892 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560070403
- Primary Citation Related Structures:
1SWF, 1SWG - PubMed Abstract:
A circularly permuted streptavidin (CP51/46) has been designed to remove the flexible polypeptide loop that undergoes an open to closed conformational change when biotin is bound. The original termini have been joined by a tetrapeptide linker, and four loop residues have been removed, resulting in the creation of new N- and C-termini. Isothermal titration calorimetric studies show that the association constant has been reduced approximately six orders of magnitude below that of wild-type streptavidin to 10(7) M(-1). The deltaH degrees of biotin association for CP51/46 is reduced by 11.1 kcal/mol. Crystal structures of CP51/46 and its biotin complex show no significant alterations in the binding site upon removal of the loop. A hydrogen bond between Ser45 and Ser52 found in the absence of biotin is broken in the closed conformation as the side-chain hydroxyl of Ser45 moves to hydrogen bond to a ureido nitrogen of biotin. This is true in both the wild-type and CP51/46 forms of the protein, and the hydrogen bonding interaction might thus help nucleate closure of the loop. The reduced entropic cost of binding biotin to CP51/46 is consistent with the removal of this loop and a reduction in entropic costs associated with loop closure and immobilization. The reduced enthalpic contribution to the free energy of binding is not readily explainable in terms of the molecular structure, as the binding contacts are nearly entirely conserved, and only small differences in solvent accessible surfaces are observed relative to wild-type streptavidin.
- Department of Bioengineering, University of Washington, Seattle 98195-7962, USA.
Organizational Affiliation: