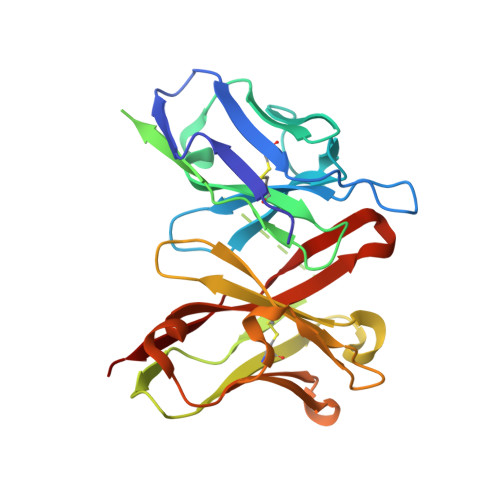
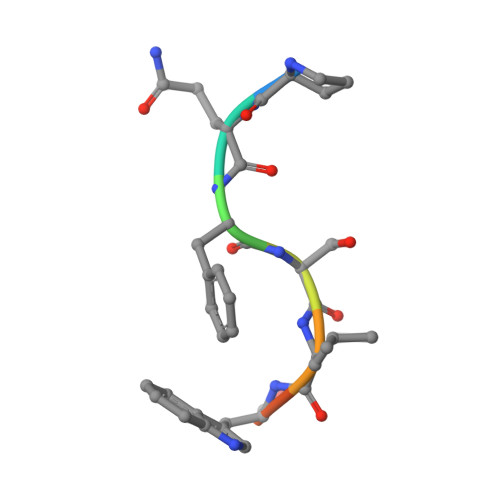
Crystal structure of a cross-reaction complex between an anti-HIV-1 protease antibody and an HIV-2 protease peptide
Rezacova, P., Brynda, J., Lescar, J., Fabry, M., Horejsi, M., Sieglova, I., Sedlacek, J., Bentley, G.A.(2005) J Struct Biol 149: 332-337
- PubMed: 15721587 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2004.11.009
- Primary Citation Related Structures:
1SVZ - PubMed Abstract:
The monoclonal antibody 1696, elicited by HIV-1 protease, inhibits the activity of both HIV-1 and HIV-2 proteases with inhibition constants in the low nanomolar range. The antibody cross-reacts with peptides derived from the N-terminal region of both proteases. The crystal structure of the recombinant single-chain Fv fragment of 1696 complexed with an N-terminal peptide from the HIV-2 protease has been determined at 1.88A resolution. Interactions of the peptide with scFv1696 are compared with the previously reported structure of scFv1696 in complex with the corresponding peptide from HIV-1 protease. The origin of cross-reactivity of mAb1696 with HIV proteases is discussed.
- Department of Recombinant Expression and Structural Biology, Institute of Molecular Genetics, Academy of Sciences of the Czech Republic, Flemingovo nam. 2, 166 37 Prague 6, Czech Republic. rezacova@img.cas.nz
Organizational Affiliation: