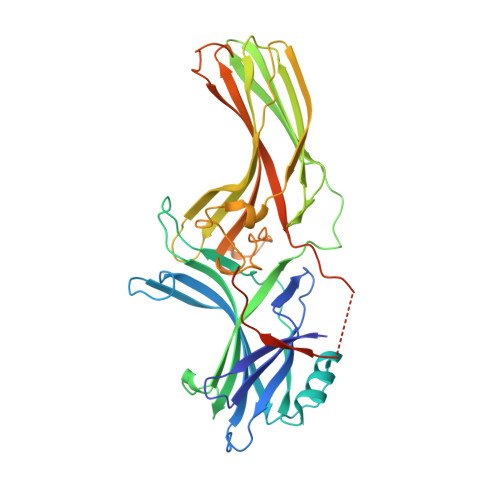
Crystal structure of cone arrestin at 2.3A: evolution of receptor specificity.
Sutton, R.B., Vishnivetskiy, S.A., Robert, J., Hanson, S.M., Raman, D., Knox, B.E., Kono, M., Navarro, J., Gurevich, V.V.(2005) J Mol Biology 354: 1069-1080
- PubMed: 16289201 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.10.023
- Primary Citation Related Structures:
1SUJ - PubMed Abstract:
Arrestins play a fundamental role in the regulation and signal transduction of G protein-coupled receptors. Here we describe the crystal structure of cone arrestin at 2.3A resolution. The overall structure of cone visual arrestin is similar to the crystal structures of rod visual and the non-visual arrestin-2, consisting of two domains, each containing ten beta-sheets. However, at the tertiary structure level, there are two major differences, in particular on the concave surfaces of the two domains implicated in receptor binding and in the loop between beta-strands I and II. Functional analysis shows that cone arrestin, in sharp contrast to its rod counterpart, bound cone pigments and non-visual receptors. Conversely, non-visual arrestin-2 bound cone pigments, suggesting that it may also regulate phototransduction and/or photopigment trafficking in cone photoreceptors. These findings indicate that cone arrestin displays structural and functional features intermediate between the specialized rod arrestin and the non-visual arrestins, which have broad receptor specificity. A unique functional feature of cone arrestin was the low affinity for its cognate receptor, resulting in an unusually rapid dissociation of the complex. Transient arrestin binding to the photopigment in cones may be responsible for the extremely rapid regeneration and reuse of the photopigment that is essential for cone function at high levels of illumination.
- Department of Neuroscience and Cell Biology, University of Texas Medical Branch, and Sealy Center for Molecular Science & Structural Biology, Galveston, TX 77555, USA.
Organizational Affiliation: