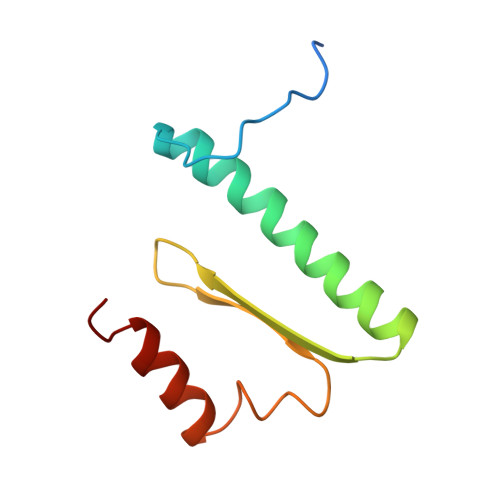
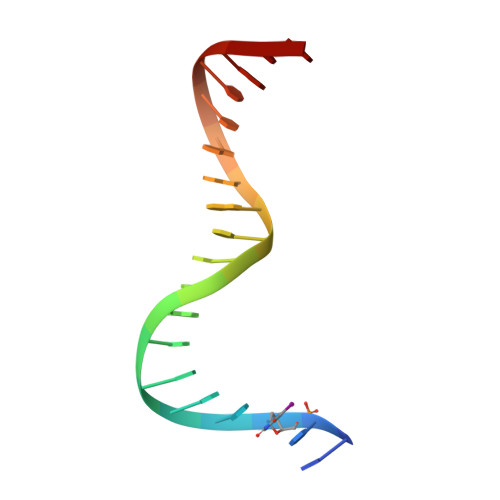
Structure of serum response factor core bound to DNA.
Pellegrini, L., Tan, S., Richmond, T.J.(1995) Nature 376: 490-498
- PubMed: 7637780 Search on PubMed
- DOI: https://doi.org/10.1038/376490a0
- Primary Citation Related Structures:
1SRS - PubMed Abstract:
The human serum response factor is a transcription factor belonging to the MADS domain protein family with members characterized from the plant and animal kingdoms. The X-ray crystal structure of the serum response factor core in a specific-recognition DNA complex shows that the functions of DNA binding, dimerization and accessory-factor interaction are compactly integrated into a novel protein unit. The intrinsic and induced conformation of the serum response element DNA is the principal DNA feature recognized in the specific complex.
- Institut für Molekularbiologie und Biophysik, ETH-Hönggerberg, Zürich, Switzerland.
Organizational Affiliation: