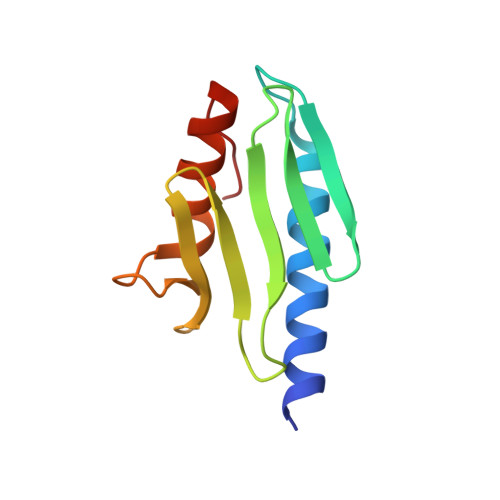
Solution Structure of the Bacterial Frataxin Ortholog, CyaY; Mapping the Iron Binding Sites
Nair, M., Adinolfi, S., Pastore, C., Kelly, G., Temussi, P., Pastore, A.(2004) Structure 12: 2037-2048
- PubMed: 15530368
- DOI: https://doi.org/10.1016/j.str.2004.08.012
- Primary Citation of Related Structures:
1SOY - PubMed Abstract:
CyaY is the bacterial ortholog of frataxin, a small mitochondrial iron binding protein thought to be involved in iron sulphur cluster formation. Loss of frataxin function leads to the neurodegenerative disorder Friedreich's ataxia. We have solved the solution structure of CyaY and used the structural information to map iron binding onto the protein surface. Comparison of the behavior of wild-type CyaY with that of a mutant indicates that specific binding with a defined stoichiometry does not require aggregation and that the main binding site, which hosts both Fe(2+) and Fe(3+), occupies a highly anionic surface of the molecule. This function is conserved across species since the corresponding region of human frataxin is also able to bind iron, albeit with weaker affinity. The presence of secondary binding sites on CyaY, but not on frataxin, hints at a possible polymerization mechanism. We suggest mutations that may provide further insights into the frataxin function.
- Division of Molecular Structure, The Ridgeway, London, NW7 1AA, United Kingdom.
Organizational Affiliation: