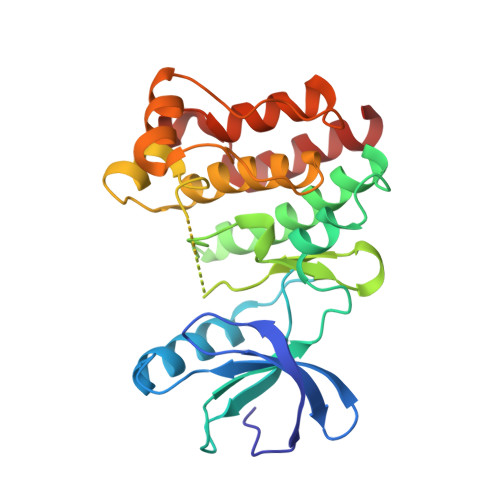
Crystal structures of interleukin-2 tyrosine kinase and their implications for the design of selective inhibitors.
Brown, K., Long, J.M., Vial, S.C., Dedi, N., Dunster, N.J., Renwick, S.B., Tanner, A.J., Frantz, J.D., Fleming, M.A., Cheetham, G.M.(2004) J Biological Chem 279: 18727-18732
- PubMed: 14766749 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M400031200
- Primary Citation Related Structures:
1SM2, 1SNU, 1SNX - PubMed Abstract:
Interleukin-2 tyrosine kinase, Itk, is an important member of the Tec family of non-receptor tyrosine kinases that play a central role in signaling through antigen receptors such as the T-cell receptor, B-cell receptor, and Fcepsilon. Selective inhibition of Itk may be an important way of modulating many diseases involving heightened or inappropriate activation of the immune system. In addition to an unliganded nonphophorylated Itk catalytic kinase domain, we determined the crystal structures of the phosphorylated and nonphosphorylated kinase domain bound to staurosporine, a potent broad-spectrum kinase inhibitor. These structures are useful for the design of novel, highly potent and selective Itk inhibitors and provide insight into the influence of inhibitor binding and phosphorylation on the conformation of Itk.
- Vertex Pharmaceuticals (Europe) Ltd., 88 Milton Park, Abingdon, Oxfordshire OX14 4RY, United Kingdom.
Organizational Affiliation: