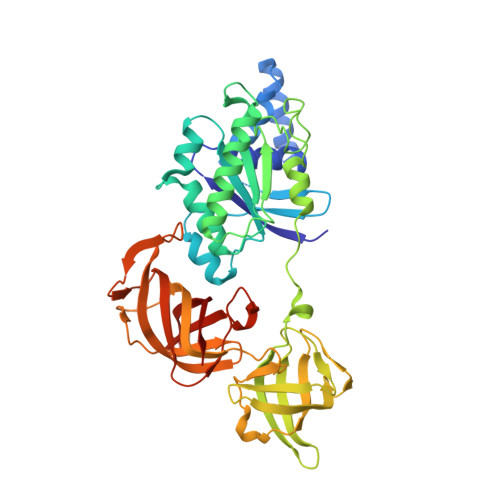
The crystal structure of Sulfolobus solfataricus elongation factor 1alpha in complex with magnesium and GDP.
Vitagliano, L., Ruggiero, A., Masullo, M., Cantiello, P., Arcari, P., Zagari, A.(2004) Biochemistry 43: 6630-6636
- PubMed: 15157096 Search on PubMed
- DOI: https://doi.org/10.1021/bi0363331
- Primary Citation Related Structures:
1SKQ - PubMed Abstract:
Recent studies have shown that elongation factors extracted from archaea/eukarya and from eubacteria exhibit different structural and functional properties. Along this line, it has been demonstrated that, in contrast to EF-Tu, Sulfolobus solfataricus EF-1alpha in complex with GDP (SsEF-1alpha.GDP) does not bind Mg(2+), when the ion is present in the crystallization medium at moderate concentration (5 mM). To further investigate the role that magnesium plays in the exchange process of EF-1alpha and to check the ability of SsEF-1alpha.GDP to bind the ion, we have determined the crystal structure of SsEF-1alpha.GDP in the presence of a nonphysiological concentration (100 mM) of Mg(2+). The analysis of the coordination of Mg(2+) unveils the structural bases for the marginal role played by the ion in the nucleotide exchange process. Furthermore, nucleotide exchange experiments carried out on a truncated form of SsEF-1alpha, consisting only of the nucleotide binding domain, demonstrate that the low affinity of SsEF-1alpha.GDP for Mg(2+) is due to the local architecture of the active site and does not depend on the presence of the other two domains. Finally, considering the available structures of EF-1alpha, a detailed mechanism for the nucleotide exchange process has been traced. Notably, this mechanism involves residues such as His14, Arg95, Gln131, and Glu134, which are strictly conserved in all archaea and eukarya EF-1alpha sequences hitherto reported.
- Istituto di Biostrutture e Bioimmagini, CNR, via Mezzocannone 6, I-80134 Napoli, Italy.
Organizational Affiliation: