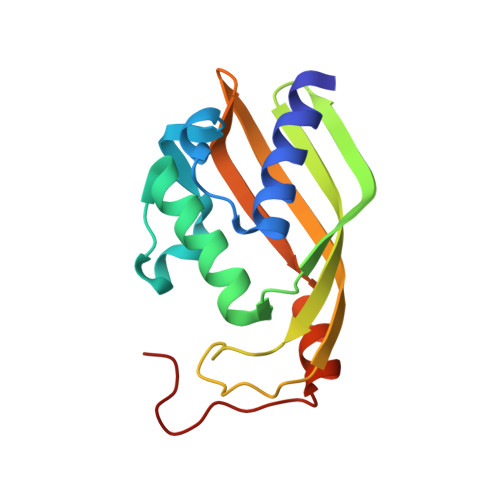
Structure of the polyketide cyclase SnoaL reveals a novel mechanism for enzymatic aldol condensation.
Sultana, A., Kallio, P., Jansson, A., Wang, J.S., Niemi, J., Schneider, G.(2004) EMBO J 23: 1911-1921
- PubMed: 15071504 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600201
- Primary Citation Related Structures:
1SJW - PubMed Abstract:
SnoaL belongs to a family of small polyketide cyclases, which catalyse ring closure steps in the biosynthesis of polyketide antibiotics produced in Streptomyces. Several of these antibiotics are among the most used anti-cancer drugs currently in use. The crystal structure of SnoaL, involved in nogalamycin biosynthesis, with a bound product, has been determined to 1.35 A resolution. The fold of the subunit can be described as a distorted alpha+beta barrel, and the ligand is bound in the hydrophobic interior of the barrel. The 3D structure and site-directed mutagenesis experiments reveal that the mechanism of the intramolecular aldol condensation catalysed by SnoaL is different from that of the classical aldolases, which employ covalent Schiff base formation or a metal ion cofactor. The invariant residue Asp121 acts as an acid/base catalyst during the reaction. Stabilisation of the enol(ate) intermediate is mainly achieved by the delocalisation of the electron pair over the extended pi system of the substrate. These polyketide cyclases thus form of family of enzymes with a unique catalytic strategy for aldol condensation.
- Department of Medical Biochemistry and Biophysics, Karolinska Institutet, Stockholm, Sweden.
Organizational Affiliation: