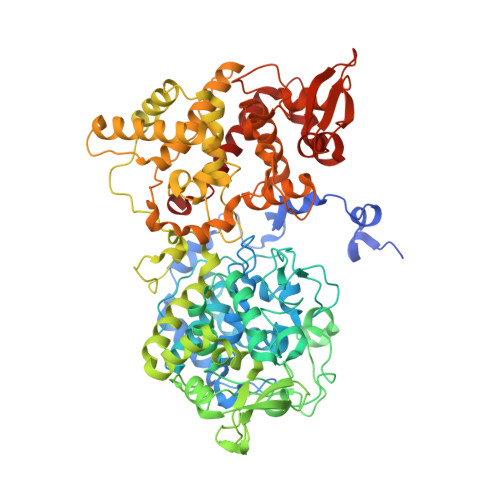
Crystal Structure of Mycobacterium tuberculosis Catalase-Peroxidase.
Bertrand, T., Eady, N.A.J., Jones, J.N., Jesmin, Nagy, J.M., Jamart-Gregoire, B., Raven, E.L., Brown, K.A.(2004) J Biological Chem 279: 38991-38999
- PubMed: 15231843 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M402382200
- Primary Citation Related Structures:
1SJ2 - PubMed Abstract:
The Mycobacterium tuberculosis catalase-peroxidase is a multifunctional heme-dependent enzyme that activates the core anti-tuberculosis drug isoniazid. Numerous studies have been undertaken to elucidate the enzyme-dependent mechanism of isoniazid activation, and it is well documented that mutations that reduce activity or inactivate the catalase-peroxidase lead to increased levels of isoniazid resistance in M. tuberculosis. Interpretation of the catalytic activities and the effects of mutations upon the action of the enzyme to date have been limited due to the lack of a three-dimensional structure for this enzyme. In order to provide a more accurate model of the three-dimensional structure of the M. tuberculosis catalase-peroxidase, we have crystallized the enzyme and now report its crystal structure refined to 2.4-A resolution. The structure reveals new information about dimer assembly and provides information about the location of residues that may play a role in catalysis including candidates for protein-based radical formation. Modeling and computational studies suggest that the binding site for isoniazid is located near the delta-meso heme edge rather than in a surface loop structure as currently proposed. The availability of a crystal structure for the M. tuberculosis catalase-peroxidase also permits structural and functional effects of mutations implicated in causing elevated levels of isoniazid resistance in clinical isolates to be interpreted with improved confidence.
- Department of Biological Sciences, Centre for Molecular Microbiology and Infection, Flowers Building, Imperial College London, London SW7 2AZ, United Kingdom.
Organizational Affiliation: