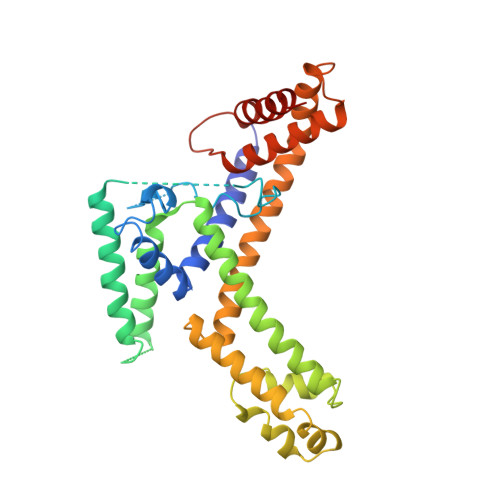
Crystal structure of a sigma 70 subunit fragment from E. coli RNA polymerase.
Malhotra, A., Severinova, E., Darst, S.A.(1996) Cell 87: 127-136
- PubMed: 8858155 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81329-x
- Primary Citation Related Structures:
1SIG - PubMed Abstract:
The 2.6 A crystal structure of a fragment of the sigma 70 promoter specificity subunit of E. coli RNA polymerase is described. Residues involved in core RNA polymerase binding lie on one face of the structure. On the opposite face, aligned along one helix, are exposed residues that interact with the -10 consensus promoter element (the Pribnow box), including four aromatic residues involved in promoter melting. The structure suggests one way in which DNA interactions may be inhibited in the absence of RNA polymerase and provides a framework for the interpretation of a large number of genetic and biochemical analyses.
- Rockefeller University, New York, New York 10021, USA.
Organizational Affiliation: