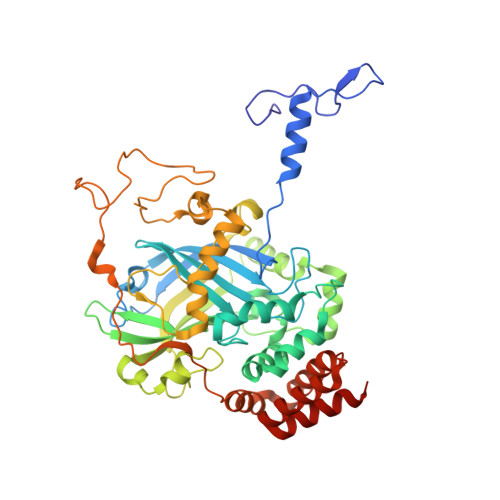
The three-dimensional structure of catalase from Enterococcus faecalis.
Hakansson, K.O., Brugna, M., Tasse, L.(2004) Acta Crystallogr D Biol Crystallogr 60: 1374-1380
- PubMed: 15272159 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904012004
- Primary Citation Related Structures:
1SI8 - PubMed Abstract:
Enterococcus faecalis haem catalase was crystallized using lithium sulfate at neutral pH. The crystals belong to space group R3, with unit-cell parameters a = b = 236.9, c = 198.1 A. The three-dimensional structure was determined by molecular replacement using a subunit of the Proteus mirabilis catalase structure. It was refined against 2.3 A synchrotron data to a free R factor of 21.8%. Like other catalases, the E. faecalis catalase is a homotetramer with a fold and structure similar to those of its structurally closest relative P. mirabilis. The solvent structure in the active site is identical in the four subunits but differs from that found in other catalases. The structural consequences of the Ramachandran outlier Ser196 are discussed.
- August Krogh Institute, Copenhagen University, Universitetsparken 13, DK-2100 Kbh O, Denmark. kohakansson@aki.ku.dk
Organizational Affiliation: