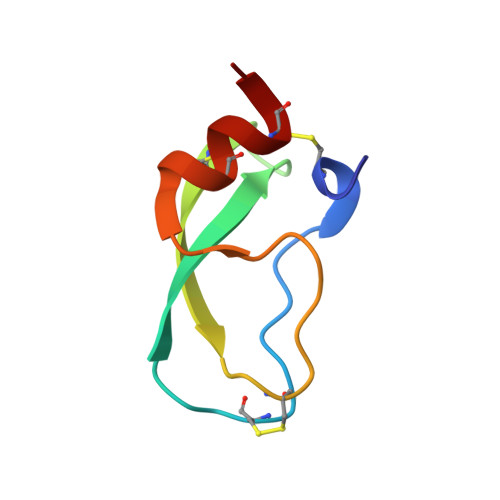
The NMR solution structure of a Kunitz-type proteinase inhibitor from the sea anemone Stichodactyla helianthus.
Antuch, W., Berndt, K.D., Chavez, M.A., Delfin, J., Wuthrich, K.(1993) Eur J Biochem 212: 675-684
- PubMed: 8462542 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.1993.tb17705.x
- Primary Citation Related Structures:
1SHP - PubMed Abstract:
The solution structure of a 55-amino-acid Kunitz-type proteinase inhibitor, ShPI, purified from the Caribbean sea anemone Stichodactyla helianthus, was determined by NMR spectroscopy. Nearly complete sequence-specific 1H-NMR assignments were obtained at pH 4.6 and 36 degrees C, and stereo-specific assignments were determined for 23 pairs of diastereotopic substituents. A data set of 666 upper distance limit constraints and 122 dihedral angle constraints collected on this basis was used as input for a structure calculation with the program DIANA. Following energy minimization with the program OPAL, the average root-mean-square diviation (RMSD) of the 20 DIANA conformers used to represent the solution structure relative to the mean structure is 61 pm for all backbone atoms N, C alpha and C', and 106 pm for all heavy atoms of residues 2-53. This high-quality solution structure of ShPI has a nearly identical molecular architecture as the bovine pancreatic trypsin inhibitor (BPTI), despite a mere 35% of sequence similarity between the two proteins. Exchange rates measured for 48 out of the 51 backbone amide protons showed that the positions of 20 slowly exchanging amide protons correlate well with hydrogen bonds involving these protons in the energy-minimized solution structure. The solution structure of ShPI is compared to the four homologous proteins for which the three-dimensional structure is also available.
- Institut für Molekularbiologie and Biophysik, Eidgenössische Technische Hochschule-Hönggerberg, Zürich, Switzerland.
Organizational Affiliation: