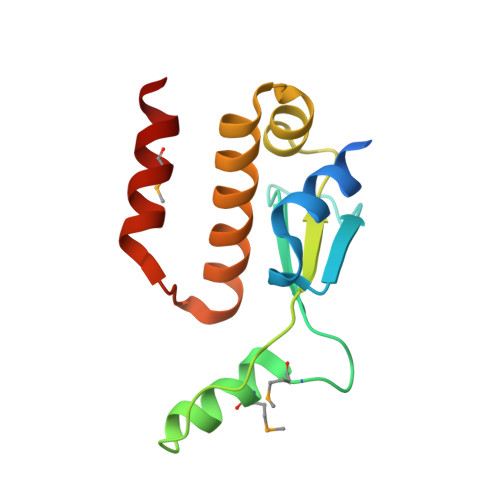
The crystal structure of the carboxy-terminal dimerization domain of htpG, the Escherichia coli Hsp90, reveals a potential substrate binding site.
Harris, S.F., Shiau, A.K., Agard, D.A.(2004) Structure 12: 1087-1097
- PubMed: 15274928 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.03.020
- Primary Citation Related Structures:
1SF8 - PubMed Abstract:
Hsp90 is a ubiquitous, well-conserved molecular chaperone involved in the folding and stabilization of diverse proteins. Beyond its capacity for general protein folding, Hsp90 influences a wide array of cellular signaling pathways that underlie key biological and disease processes. It has been proposed that Hsp90 functions as a molecular clamp, dimerizing through its carboxy-terminal domain and utilizing ATP binding and hydrolysis to drive large conformational changes including transient dimerization of the amino-terminal and middle domains. We have determined the 2.6 A X-ray crystal structure of the carboxy-terminal domain of htpG, the Escherichia coli Hsp90. This structure reveals a novel fold and that dimerization is dependent upon the formation of a four-helix bundle. Remarkably, proximal to the helical dimerization motif, each monomer projects a short helix into solvent. The location, flexibility, and amphipathic character of this helix suggests that it may play a role in substrate binding and hence chaperone activity.
- Department of Biochemistry and Biophysics, University of California, San Francisco, San Francisco, CA 94143, USA.
Organizational Affiliation: