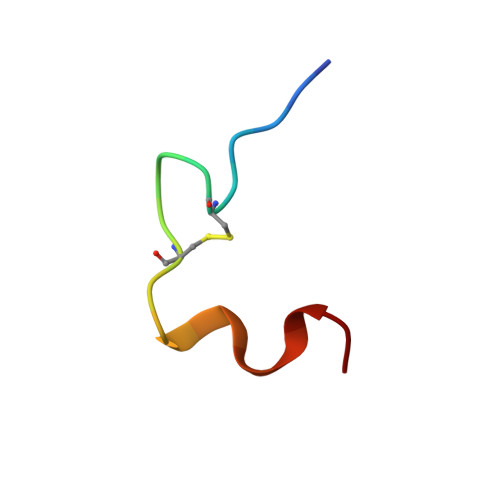
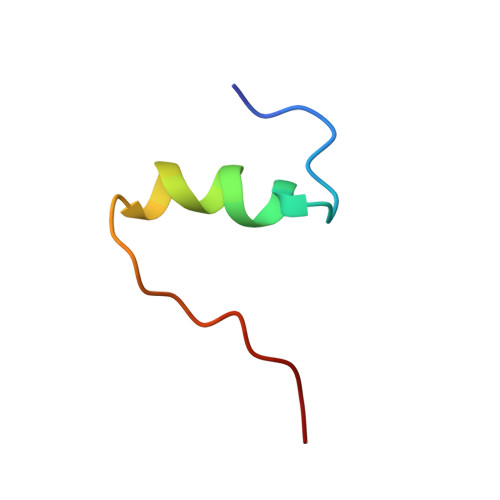
Mechanism of insulin fibrillation: the structure of insulin under amyloidogenic conditions resembles a protein-folding intermediate
Hua, Q.X., Weiss, M.A.(2004) J Biological Chem 279: 21449-21460
- PubMed: 14988398 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M314141200
- Primary Citation Related Structures:
1SF1 - PubMed Abstract:
Insulin undergoes aggregation-coupled misfolding to form a cross-beta assembly. Such fibrillation has long complicated its manufacture and use in the therapy of diabetes mellitus. Of interest as a model for disease-associated amyloids, insulin fibrillation is proposed to occur via partial unfolding of a monomeric intermediate. Here, we describe the solution structure of human insulin under amyloidogenic conditions (pH 2.4 and 60 degrees C). Use of an enhanced sensitivity cryogenic probe at high magnetic field avoids onset of fibrillation during spectral acquisition. A novel partial fold is observed in which the N-terminal segments of the A- and B-chains detach from the core. Unfolding of the N-terminal alpha-helix of the A-chain exposes a hydrophobic surface formed by native-like packing of the remaining alpha-helices. The C-terminal segment of the B-chain, although not well ordered, remains tethered to this partial helical core. We propose that detachment of N-terminal segments makes possible aberrant protein-protein interactions in an amyloidogenic nucleus. Non-cooperative unfolding of the N-terminal A-chain alpha-helix resembles that observed in models of proinsulin folding intermediates and foreshadows the extensive alpha --> beta transition characteristic of mature fibrils.
- Department of Biochemistry, Case Western Reserve University School of Medicine, Cleveland, Ohio 44016-4935.
Organizational Affiliation: