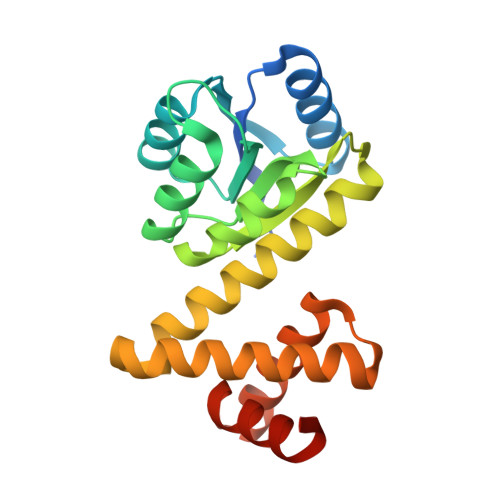
The Crystal and Solution Structure of a Putative Transcriptional Antiterminator from Mycobacterium tuberculosis.
Morth, J.P., Feng, V., Perry, L.J., Svergun, D.I., Tucker, P.A.(2004) Structure 12: 1595-1605
- PubMed: 15341725 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.06.018
- Primary Citation Related Structures:
1S8N, 1SD5 - PubMed Abstract:
We describe the crystal structure of Rv1626 from Mycobacterium tuberculosis at 1.48 A resolution and the corresponding solution structure determined from small angle X-ray scattering. The N-terminal domain shows structural homology to the receiver domains found in bacterial two-component systems. The C-terminal domain has high structural homology to a recently discovered RNA binding domain involved in transcriptional antitermination. The molecule in solution was found to be monomeric as it is in the crystal, but in solution it undergoes a conformational change that is triggered by changes in ionic strength. This is the first structure that links the phosphorylation cascade of the two-component systems with the antitermination event in the transcriptional machinery. Rv1626 belongs to a family of proteins, which we propose calling phosphorylation-dependent transcriptional antitermination regulators, so far only found in bacteria, and includes NasT, a protein from the assimilatory nitrate/nitrite reductase operon of Azetobacter vinelandii.
- EMBL Hamburg Outstation, c/o DESY, Notkestrasse 85, Germany.
Organizational Affiliation: