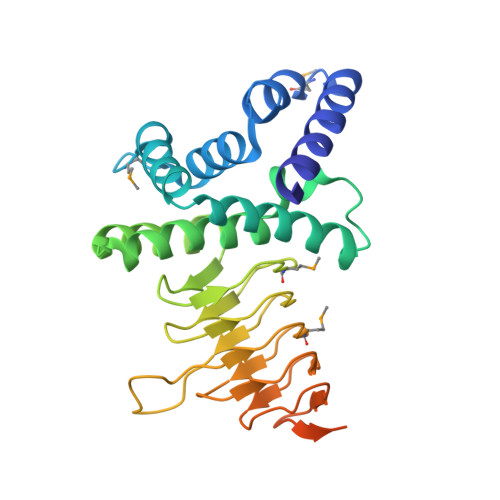
Structure of serine acetyltransferase from Haemophilus influenzae Rd.
Gorman, J., Shapiro, L.(2004) Acta Crystallogr D Biol Crystallogr 60: 1600-1605
- PubMed: 15333931 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904015240
- Primary Citation Related Structures:
1S80 - PubMed Abstract:
The crystal structure of serine acetyltransferase (SAT) from Haemophilus influenzae Rd determined at 2.7 A resolution is presented. SAT is a member of a family of hexapeptide-containing transferases that contain six-residue tandem repeats (LIV)-G-X(4) that have been shown to form left-handed parallel beta-helices. In the current structure, each protomer is comprised of two domains: an N-terminal alpha-helical domain and a C-terminal left-handed parallel beta-helix domain. Although other members of this protein family are known to form trimeric structures, SAT forms a dimer of trimers in which the trimer interface is mediated through interactions between both the beta-helix domains and N-terminal domains; these trimers dimerize through contacts in the N-terminal domain. All dimer-of-trimer interactions are mediated through amino acids within an N-terminal extension common only to a subset of SATs, suggesting that members of this subfamily may also adopt hexameric structures. Putative active sites are formed by crevices between adjacent protomers in a trimer. Thus, six independent active sites exist in the hexameric enzyme complex.
- Department of Ophthalmology, Columbia University College of Physicians and Surgeons, 630 West 168th Street, New York, NY 10032, USA.
Organizational Affiliation: