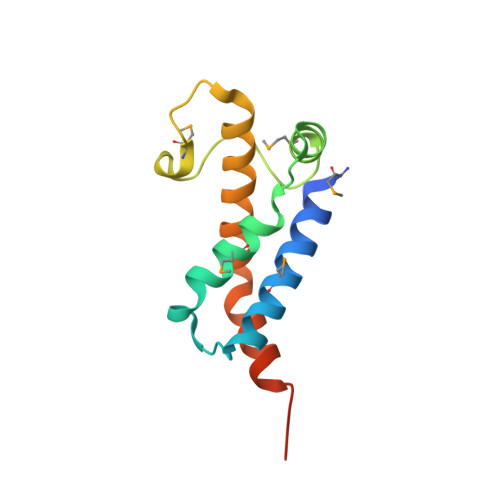
Structure of Ocr from Bacteriophage T7, a Protein that Mimics B-Form DNA
Walkinshaw, M.D., Taylor, P., Sturrock, S.S., Atanasiu, C., Berg, T., Henderson, R.M., Edwardson, J.M., Dryden, D.T.(2002) Mol Cell 9: 187-194
- PubMed: 11804597 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(02)00435-5
- Primary Citation Related Structures:
1S7Z - PubMed Abstract:
We have solved, by X-ray crystallography to a resolution of 1.8 A, the structure of a protein capable of mimicking approximately 20 base pairs of B-form DNA. This ocr protein, encoded by gene 0.3 of bacteriophage T7, mimics the size and shape of a bent DNA molecule and the arrangement of negative charges along the phosphate backbone of B-form DNA. We also demonstrate that ocr is an efficient inhibitor in vivo of all known families of the complex type I DNA restriction enzymes. Using atomic force microscopy, we have also observed that type I enzymes induce a bend in DNA of similar magnitude to the bend in the ocr molecule. This first structure of an antirestriction protein demonstrates the construction of structural mimetics of long segments of B-form DNA.
- Institute of Cell and Molecular Biology, The King's Buildings, University of Edinburgh, EH9 3JR, Edinburgh, United Kingdom. malcolm.walkinsahaw@ed.ac.uk
Organizational Affiliation: