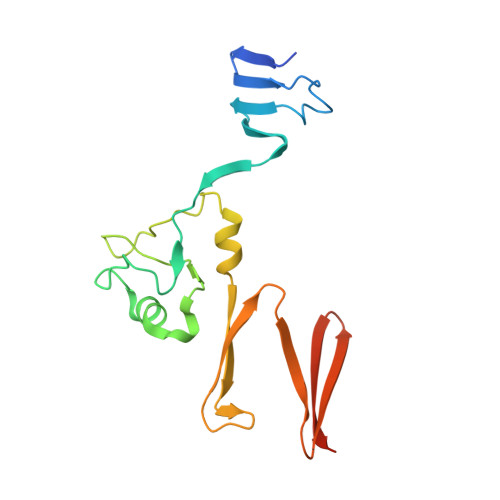
Structural basis for host recognition by the Haemophilus influenzae Hia autotransporter.
Yeo, H.J., Cotter, S.E., Laarmann, S., Juehne, T., St Geme, J.W., Waksman, G.(2004) EMBO J 23: 1245-1256
- PubMed: 15029242
- DOI: https://doi.org/10.1038/sj.emboj.7600142
- Primary Citation of Related Structures:
1S7M - PubMed Abstract:
Haemophilus influenzae is an important human pathogen that initiates infection by colonizing the upper respiratory tract. The H. influenzae Hia autotransporter is an adhesive protein that promotes adherence to respiratory epithelial cells. Hia adhesive activity resides in two homologous binding domains, called HiaBD1 and HiaBD2. These domains interact with the same host cell receptor, but bind with different affinities. In this report, we describe the crystal structure of the high-affinity HiaBD1 binding domain, which has a novel trimeric architecture with three-fold symmetry and a mushroom shape. The subunit constituents of the trimer are extensively intertwined. The receptor-binding pocket is formed by an acidic patch that is present on all three faces of the trimer, providing potential for a multivalent interaction with the host cell surface, analogous to observations with the trimeric tumor necrosis factor superfamily of proteins. Hia is a novel example of a bacterial trimeric adhesin and may be the prototype member of a large family of bacterial virulence proteins with a similar architecture.
- Institute of Structural Molecular Biology, Birkbeck and University College London, Malet Street, London, UK.
Organizational Affiliation: