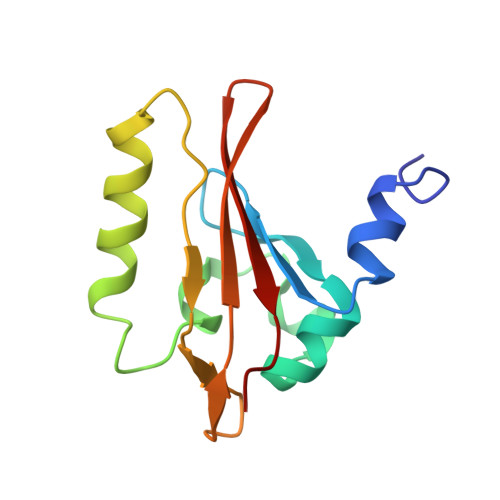
Insights into signal transduction involving PAS domain oxygen-sensing heme proteins from the X-ray crystal structure of Escherichia coli Dos heme domain (Ec DosH)
Park, H.J., Suquet, C., Satterlee, J.D., Kang, C.H.(2004) Biochemistry 43: 2738-2746
- PubMed: 15005609 Search on PubMed
- DOI: https://doi.org/10.1021/bi035980p
- Primary Citation Related Structures:
1S66, 1S67 - PubMed Abstract:
The X-ray crystal structure of the Escherichia coli (Ec) direct oxygen sensor heme domain (Ec DosH) has been solved to 1.8 A using Fe multiple-wavelength anomalous dispersion (MAD), and the positions of Met95 have been confirmed by selenomethionine ((Se)Met) MAD. Ec DosH is the sensing part of a larger two-domain sensing/signaling protein, in which the signaling domain has phosphodiesterase activity. The asymmetric unit of the crystal lattice contains a dimer comprised of two differently ligated heme domain monomers. Except for the heme ligands, the monomer heme domains are identical. In one monomer, the heme is ligated by molecular oxygen (O(2)), while in the other monomer, an endogenous Met95 with S --> Fe ligation replaces the exogenous O(2) ligand. In both heme domains, the proximal ligand is His77. Analysis of these structures reveals sizable ligand-dependent conformational changes in the protein chain localized in the FG turn, the G(beta)-strand, and the HI turn. These changes provide insight to the mechanism of signal propagation within the heme domain following initiation due to O(2) dissociation.
- School of Molecular Biosciences, Washington State University, Pullman, Washington 99164-4660, USA.
Organizational Affiliation: