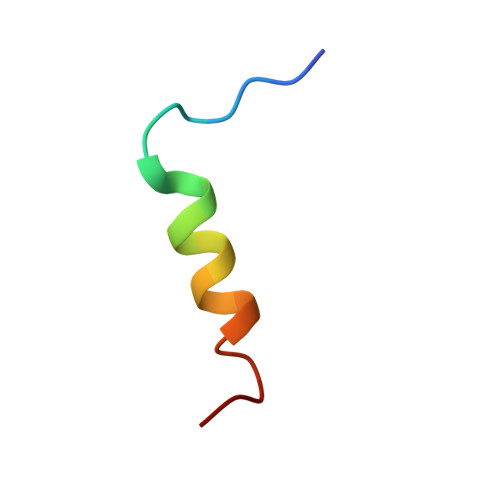
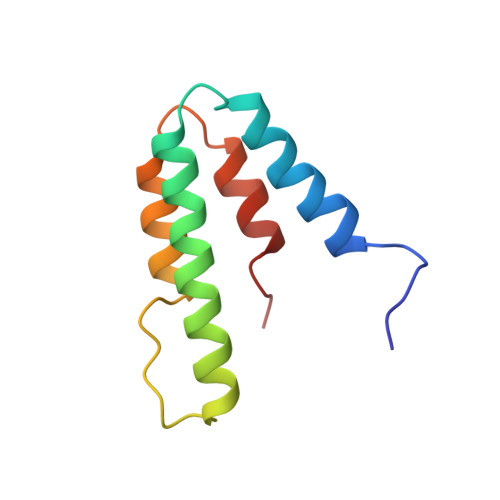
HBP1 and Mad1 repressors bind the Sin3 corepressor PAH2 domain with opposite helical orientations.
Swanson, K.A., Knoepfler, P.S., Huang, K., Kang, R.S., Cowley, S.M., Laherty, C.D., Eisenman, R.N., Radhakrishnan, I.(2004) Nat Struct Mol Biol 11: 738-746
- PubMed: 15235594
- DOI: https://doi.org/10.1038/nsmb798
- Primary Citation Related Structures:
1S5Q, 1S5R - PubMed Abstract:
Recruitment of the histone deacetylase (HDAC)-associated Sin3 corepressor is an obligatory step in many eukaryotic gene silencing pathways. Here we show that HBP1, a cell cycle inhibitor and regulator of differentiation, represses transcription in a HDAC/Sin3-dependent manner by targeting the mammalian Sin3A (mSin3A) PAH2 domain. HBP1 is unrelated to the Mad1 repressor for which high-resolution structures in complex with PAH2 have been described. We show that like Mad1, the HBP1 transrepression domain binds through a helical structure to the hydrophobic cleft of mSin3A PAH2. Notably, the HBP1 helix binds PAH2 in a reversed orientation relative to Mad1 and, equally unexpectedly, this is correlated with a chain reversal of the minimal Sin3 interaction motifs. These results not only provide insights into how multiple, unrelated transcription factors recruit the same coregulator, but also have implications for how sequence similarity searches are conducted.
- Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, Illinois 60208-3500, USA.
Organizational Affiliation: